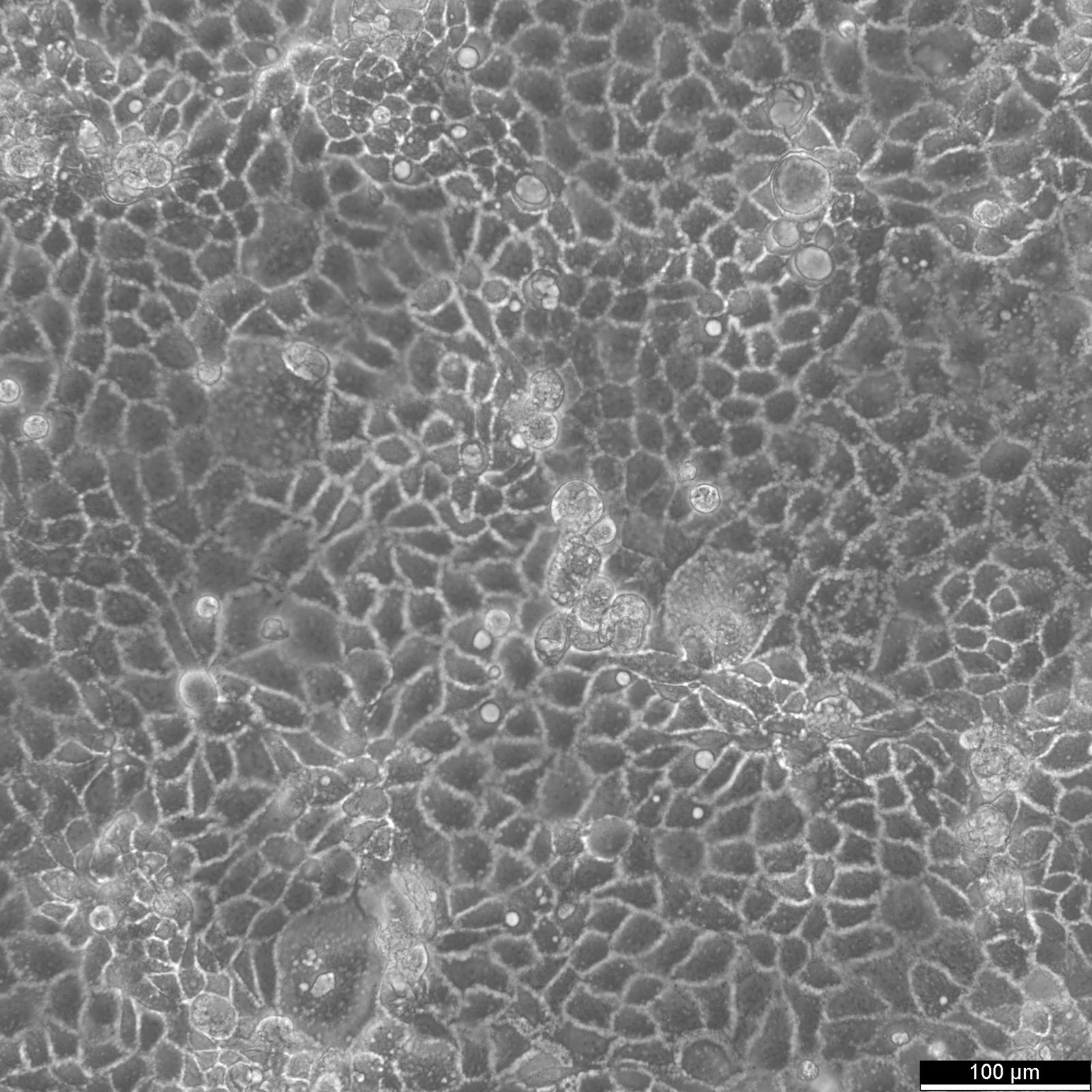
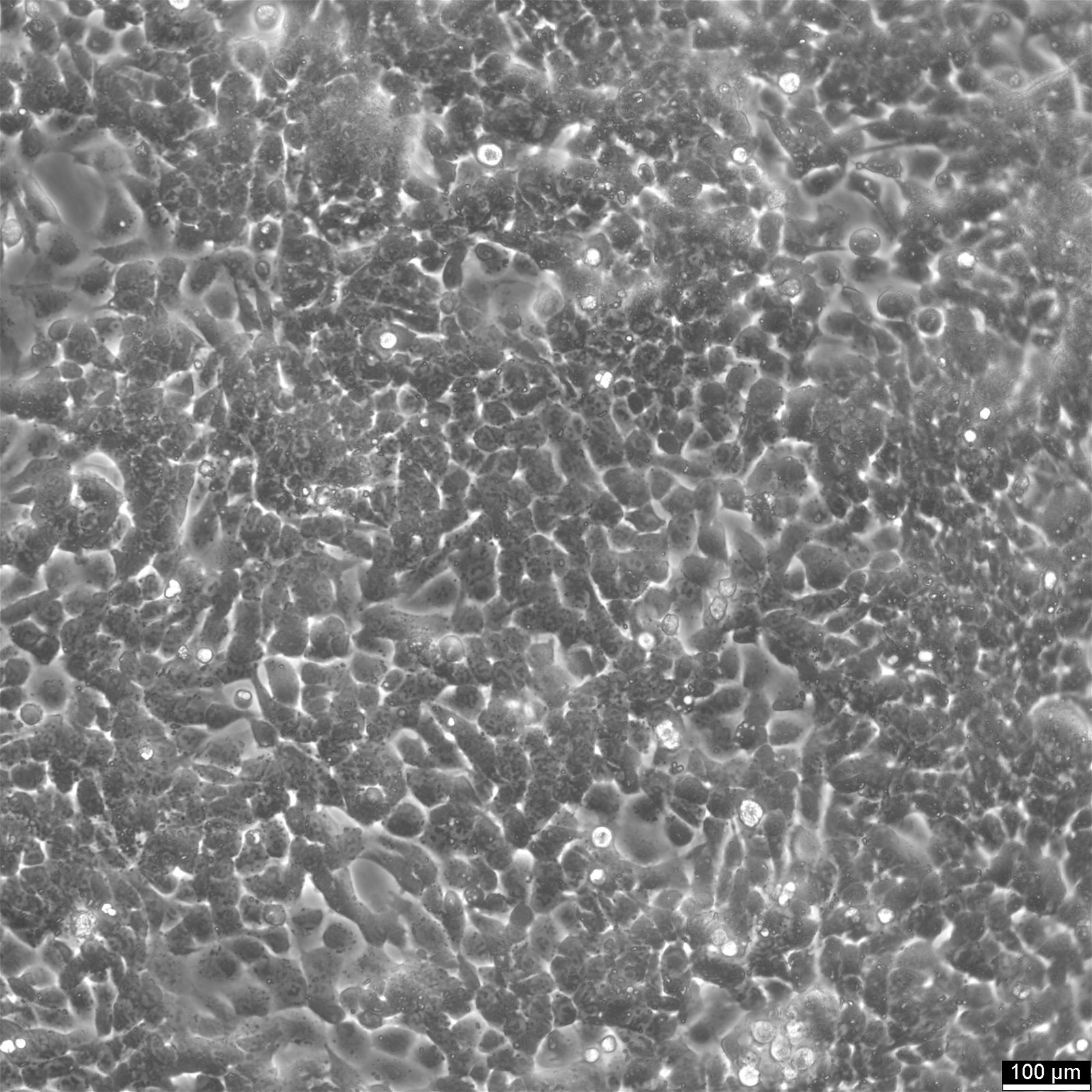
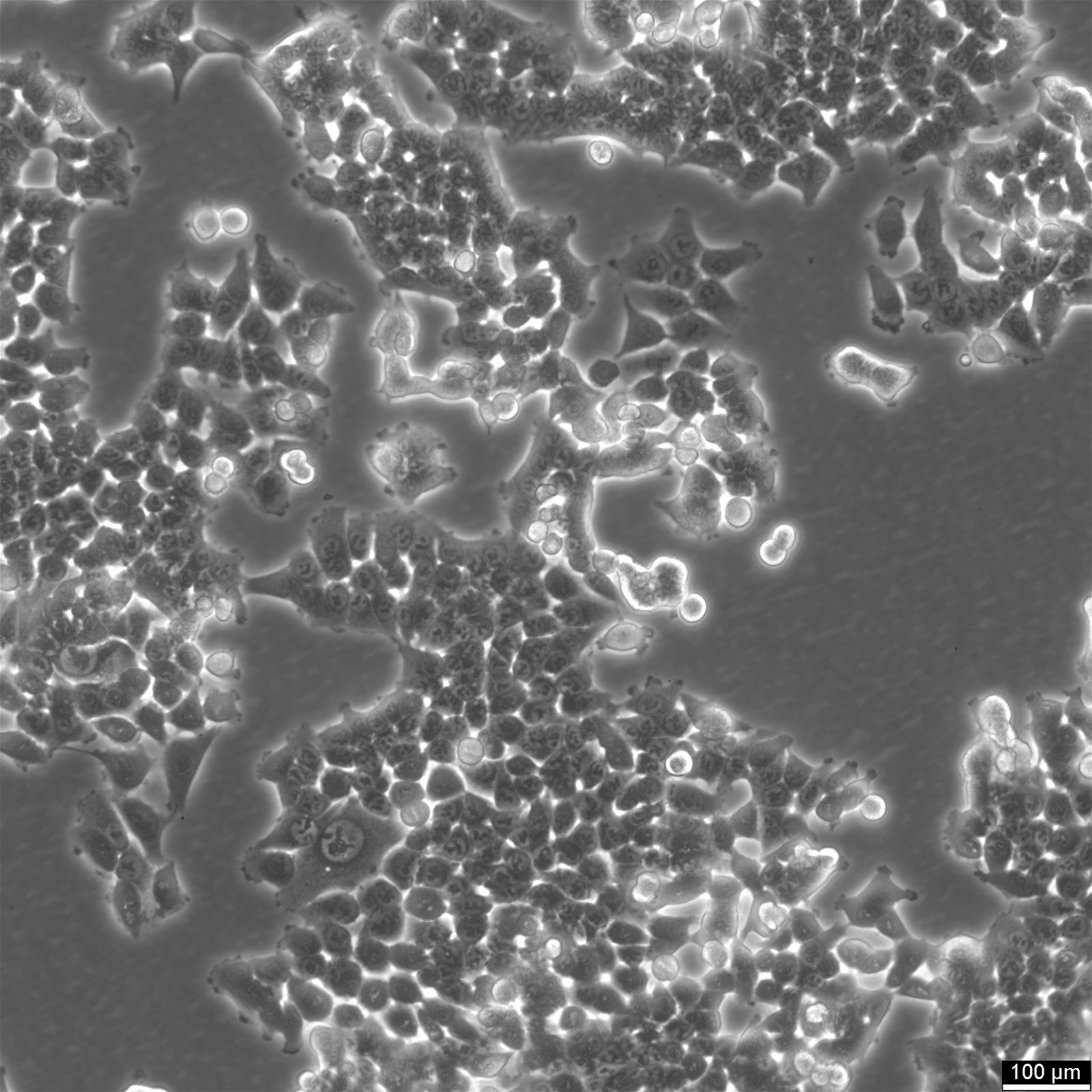
HCT116 Cells






USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
Essential facts about HCT116 cells
| Description | HCT116 cells, isolated from a colon cancer patient, serve a crucial role in therapeutic studies and drug screenings, particularly in colon cancer research. HCT-116 cells are recognized for a mutation in codon 13 of the KRAS proto-oncogene, highlighting their utility in gene therapy research, especially since they are amenable to transfection with viral vectors. In apoptosis research, HCT116 cells are pivotal for studying the mechanisms of apoptosis and cell death. The effects of butyrate, a short-chain fatty acid, have been extensively studied in HCT116 cells, revealing that butyrate inhibits colon cancer proliferation by inducing apoptosis, highlighting the intricate cancer-cell interaction and the broader implications for cancer research. Butyrate's role in modulating gene expression changes and inducing endoplasmic reticulum stress response in HCT116 cells underscores the cellular complexity in colorectal cancer cell lines. The interaction between HCT116 colon cancer cells and therapeutic agents like metformin, known for its legacy effect and potential to reduce the risk of cancer, is of significant interest. Metformin's influence on HCT116 colon cell proliferation, p21 protein level modulation, and its broader implications on proliferation and growth offer insights into the management of primary tumors and the prevention of tumors and metastases. HCT116 cells are invaluable for oncological research, providing critical insights into the efficacy of therapeutics and the molecular dynamics of cancer progression. With a notable KRAS mutation and susceptibility to transfection, these cells facilitate gene therapy studies, apoptosis analysis, and colorectal cancer treatment and prevention strategies. |
|---|---|
| Organism | Human |
| Tissue | Colorectal |
| Disease | Adenocarcinoma |
| Synonyms | HCT-116, HCT.116, HCT_116, HCT 116, CoCL2 |
Features of the colorectal carcinoma cell line HCT116
| Age | 48 years |
|---|---|
| Gender | Male |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Adherent |
Documentation
| Citation | HCT116 (Cytion catalog number 300195) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_0291 |
Genomics of HCT116 colorectal cancer cells
| Antigen expression | The cells are positive for keratin by immunoperoxidase staining. HCT 116 cells are positive for transforming growth factor beta 1 (TGF beta 1) and beta 2 (TGF beta 2) expression. |
|---|---|
| Tumorigenic | Yes, in nude mice (inoculum of 5-10 x 106 cells) |
| Ploidy status | Aneuploid |
| MSI-status | Instable (MSI-high) |
| Karyotype | The karyotype of HCT116 cells is nearly diploid, with 70% of the cells harboring 45 chromosomes, often showing an overrepresentation of chromosomes 8, 10, 16, and 17 on the long arms, along with the absence of the Y-chromosome. |
Culturing guidelines
| Culture Medium | McCoys 5a, w: 3.0 g/L Glucose, w: stable Glutamine, w: 2.0 mM Sodium pyruvate, w: 2.2 g/L NaHCO3 (Cytion article number 820200a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Doubling time | 25 to 35 hours |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 2 x 104 cells/cm2 |
| Fluid renewal | 1 to 2 times per week |
| Post-Thaw Recovery | 3 days |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Cell Purity and Identity
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300195-150523 | Certificate of Analysis | 23. May. 2025 | 300195 |
| 300195-160424 | Certificate of Analysis | 23. May. 2025 | 300195 |
-
Related products
Related products