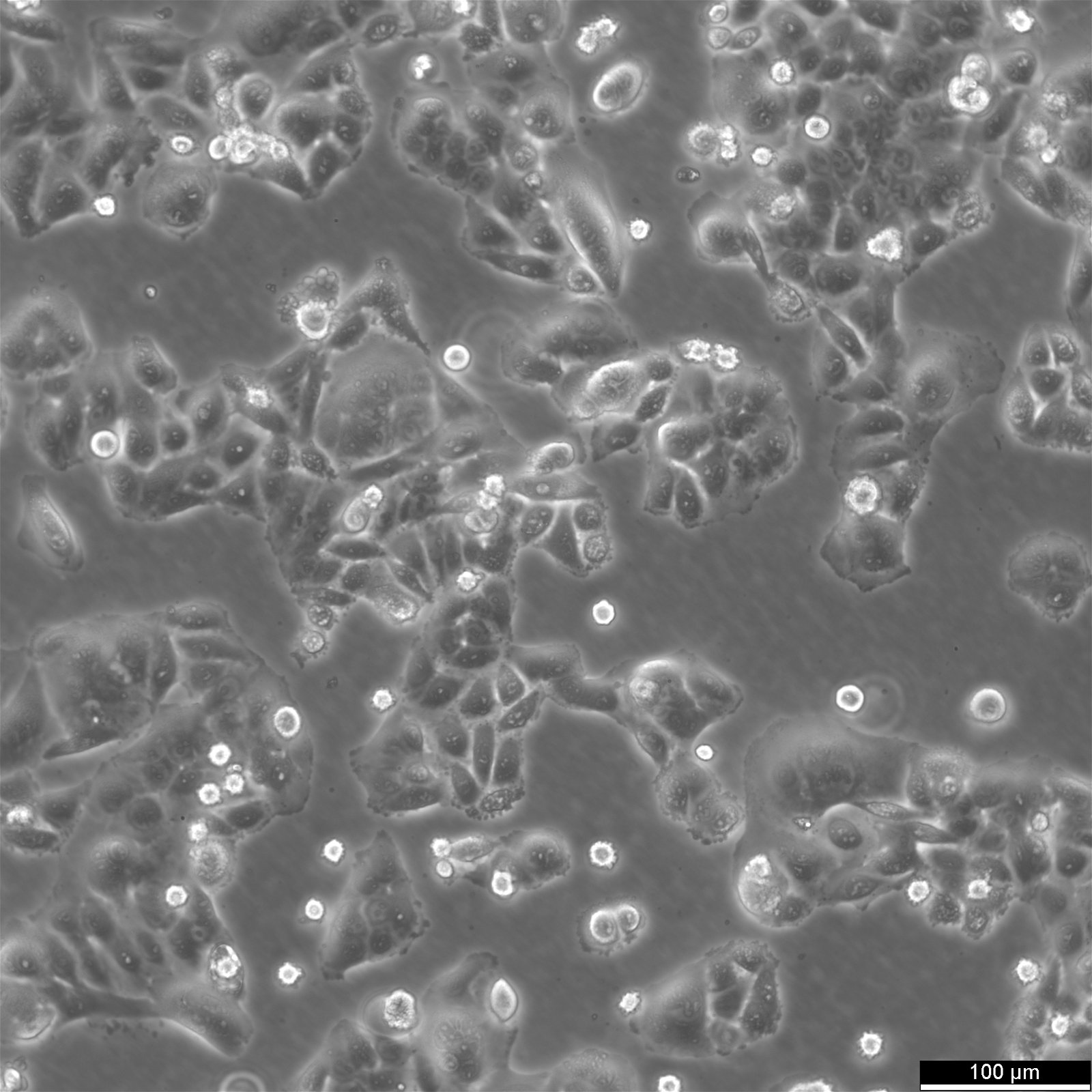
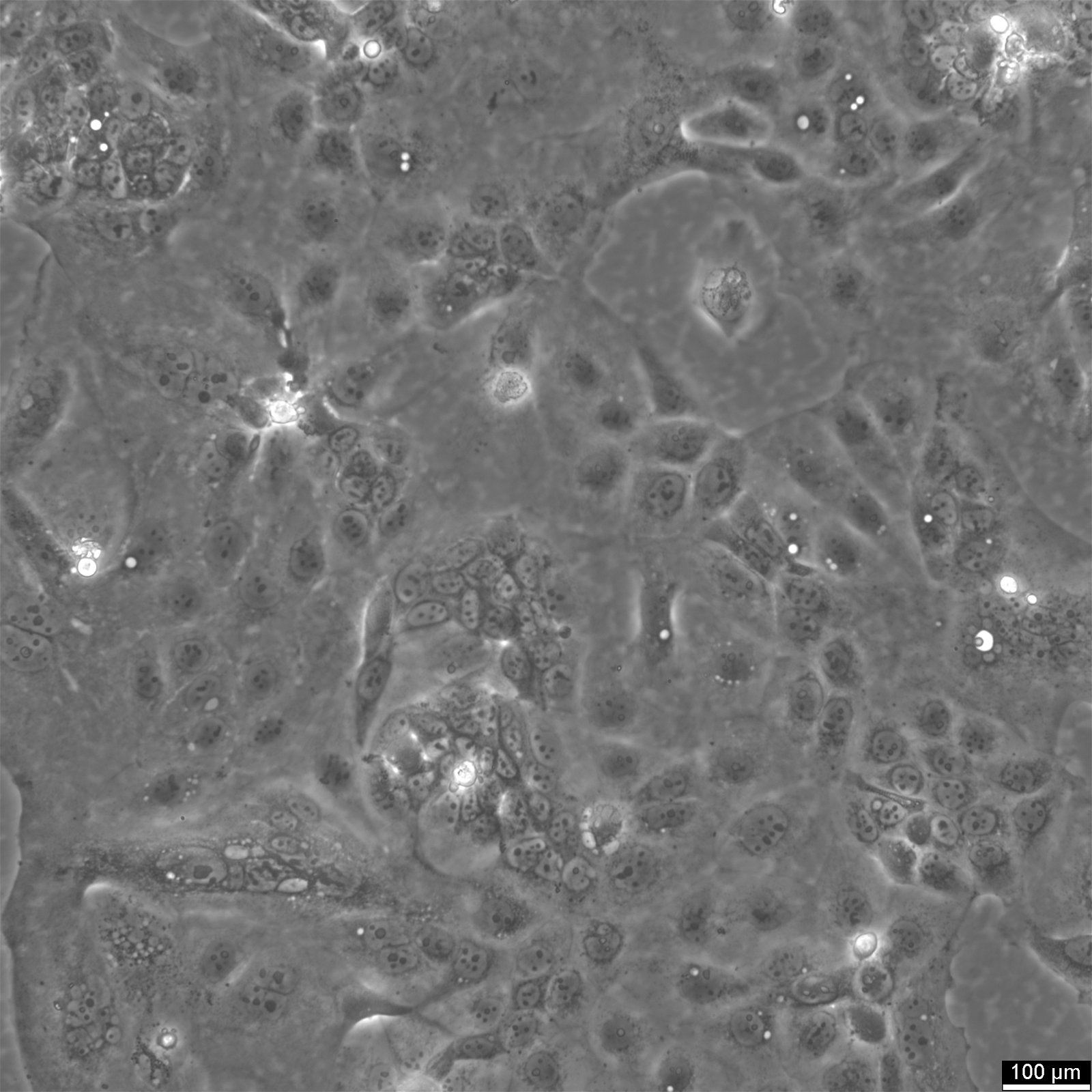
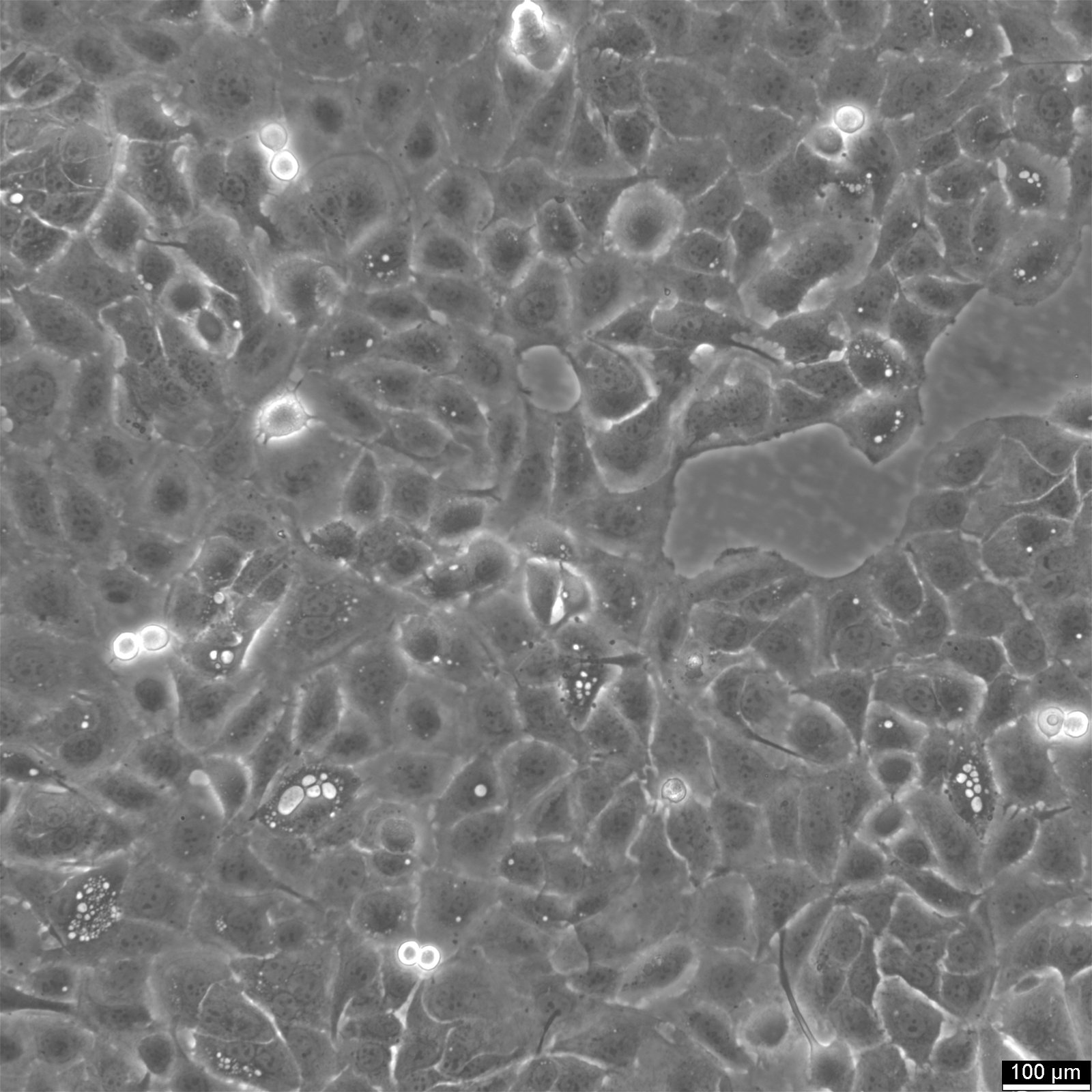
SW620 Cells
































USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
Basic details about the SW-620 cancer cell line
| Description | The SW-620 cell line, originating from the large intestine of a 51-year-old male with Dukes-C colorectal cancer, is a pivotal research model in colorectal cancer, especially for cancer biomarkers, chemotherapy, and the study of metastatic cancer cells. SW-620 cells are pivotal for studying cell apoptosis and the resistance mechanisms to anoikis, a form of programmed cell death crucial for preventing metastasis. Research utilizing the SW-620 colon cancer cells has delved into proteomic analysis to understand proteome changes under different conditions, such as hypoxia. Hypoxic SW620 cells exhibit specific proteome adaptations that contribute to chemotherapy resistance. SW620 colon cancer cells have been pivotal in evaluating natural compounds like curcumin and their impact on cancer cell viability. Studies have shown that curcumin inhibits cell viability in SW-620 cells. Moreover, the cell line aids in assessing the effects of chemotherapeutic agents and the potential for chemotherapy resistance, which is critical for advancing cancer treatment strategies. Exhibiting high tumorigenic and metastatic capabilities, SW-620 cells form solid tumors in vivo. The SW620 xenograft model, alongside the study of specific pathways like the catenin pathway and the role of transcription factors such as cdx2 in colonic adenocarcinoma cells, enriches our understanding of colorectal cancer's molecular underpinnings. In summary, SW-620 human colon adenocarcinoma cells are an invaluable resource in cancer research, offering a multifaceted approach to understanding colorectal cancer's complexities. |
|---|---|
| Organism | Human |
| Tissue | Colorectal |
| Disease | Adenocarcinoma |
| Metastatic site | Lymph node |
| Synonyms | SW620, SW 620, SW.620 |
Aspects
| Age | 51 years |
|---|---|
| Gender | Male |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Adherent |
Specification
| Citation | SW-620 (Cytion catalog number 300466) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_0547 |
Genetic makeup of the colorectal cancer cell line SW620
| Tumorigenic | Yes, in athymic nude mice |
|---|---|
| Karyotype | Average number of chromosomes 48 (range, 46-52). Eighteen marker chromosomes. For a detailed description of the karyotype we refer to Melcher et al. |
Maintenance techniques
| Culture Medium | DMEM, w: 4.5 g/L Glucose, w: 4 mM L-Glutamine, w: 3.7 g/L NaHCO3, w: 1.0 mM Sodium pyruvate (Cytion article number 820300a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Fluid renewal | 2 times per week |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality assurance of SW620 cells
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300466-322F | Certificate of Analysis | 23. May. 2025 | 300466 |
| 300466-150925 | Certificate of Analysis | 05. Dec. 2025 | 300466 |
-
Related products
Related products
LoVo Cell LineOrganism Human Tissue Colon, grade IV, Dukes' type C Disease Adenocarcinoma USD$395.00*SW480 CellsOrganism Human Tissue Colon Disease Adenocarcinoma, Grade IV, Dukes' type B. USD$395.00*