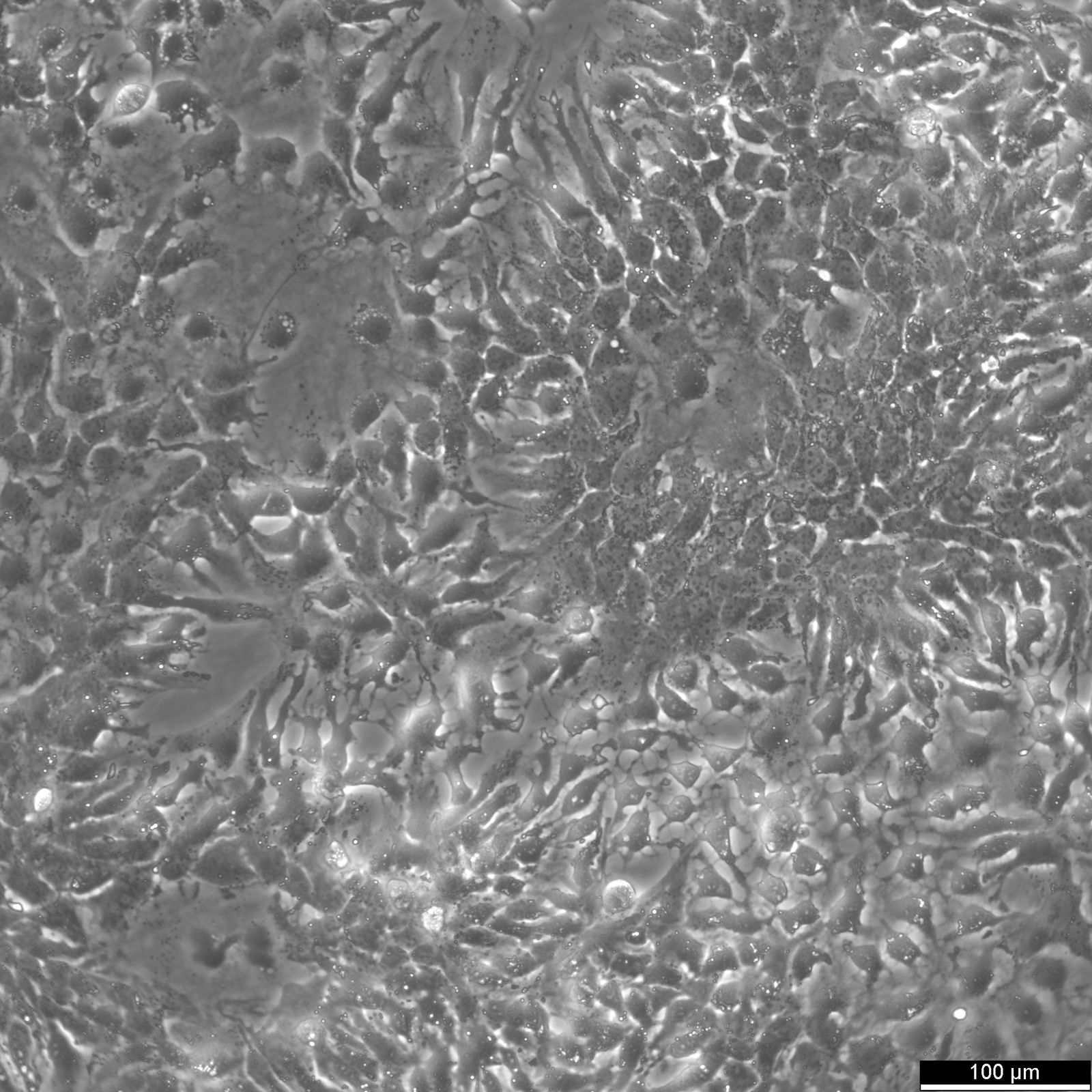
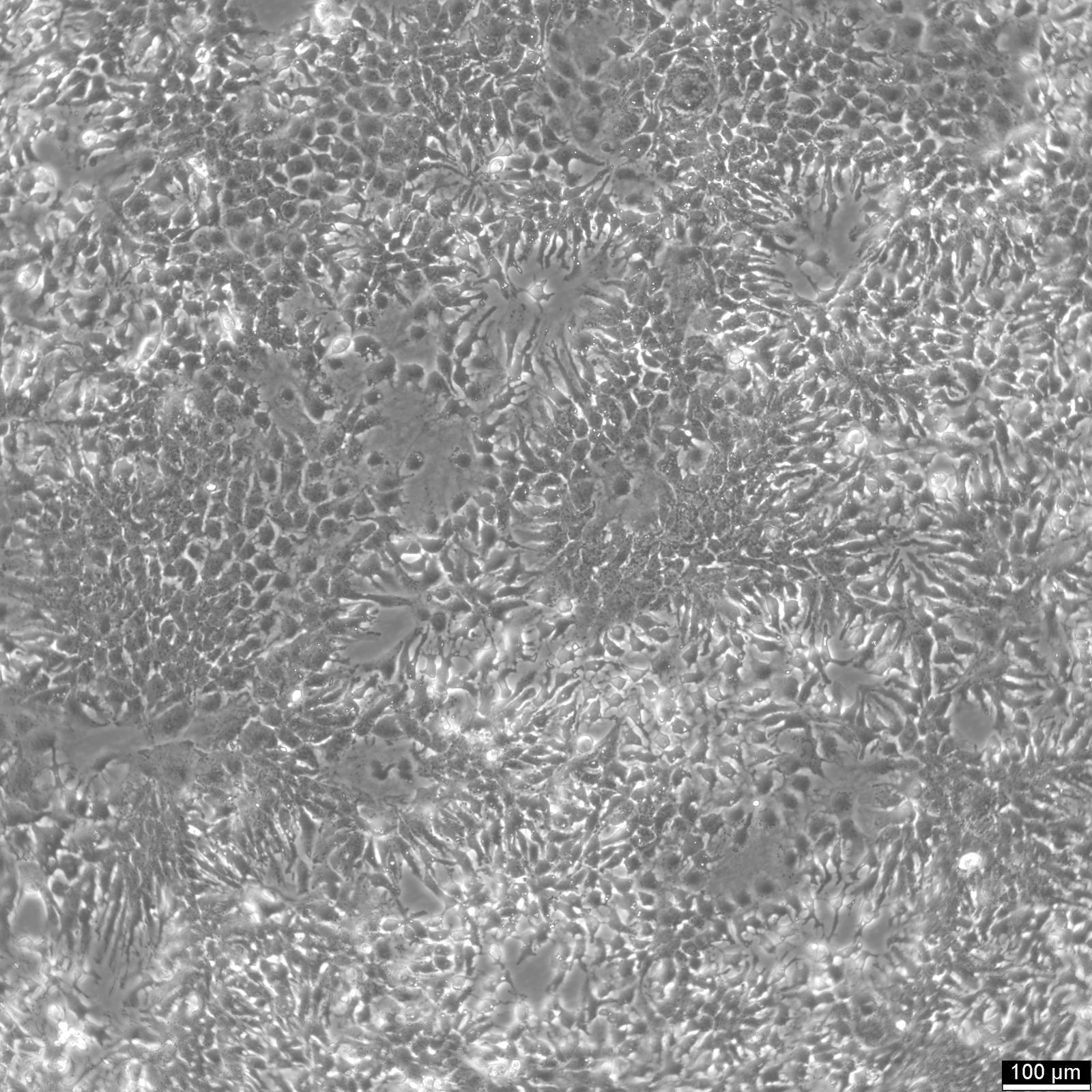
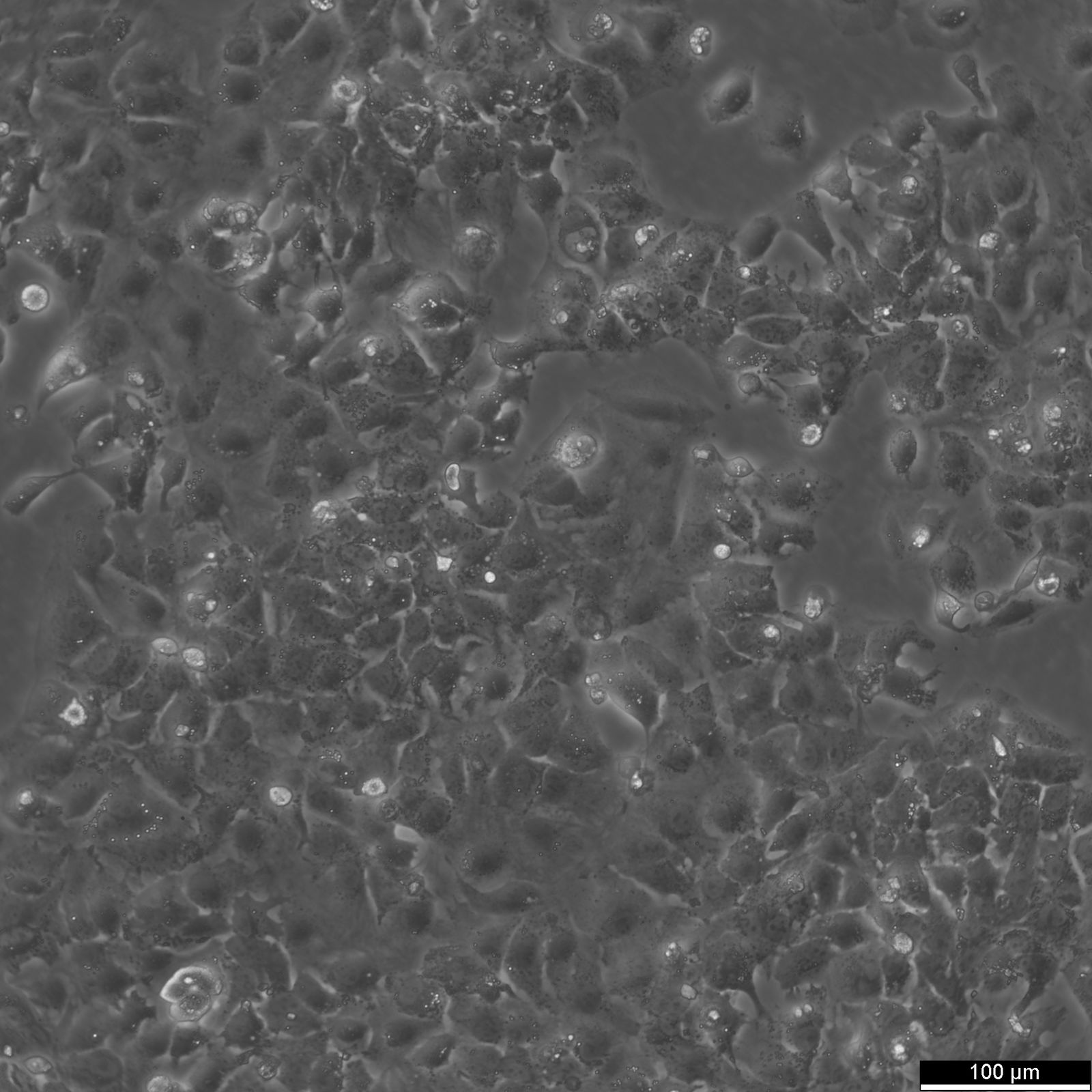
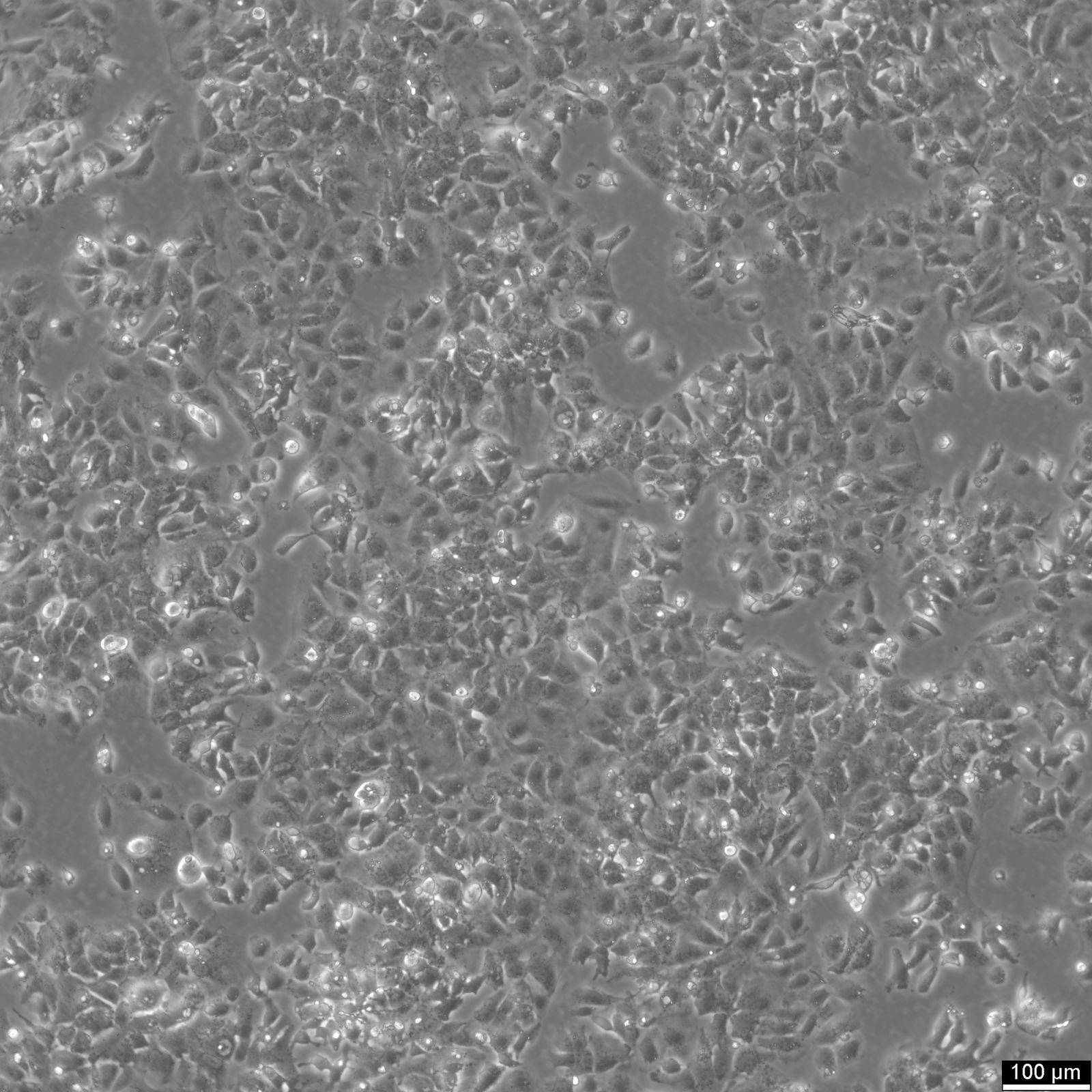
HuH7 Cells












USD$550.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
Insights on HuH-7 cell lines
| Description | HuH-7 cells are a type of epithelial-like, tumorigenic cell line initially taken from a liver tumor in a 57-year-old Japanese male in 1982. The human hepatoma-derived HuH-7 cell line and its derivatives have been widely used in research as a convenient experimental substitute for primary hepatocytes. In particular, they have been instrumental in hepatitis C research and used as host cells for propagating the virus in vitro. HuH-7 cells have played a crucial role in hepatitis C research, especially when it comes to drug development. Prior to 2005, researchers were unable to cultivate the hepatitis C virus in the laboratory, making it difficult to test potential drug candidates against it. The introduction of the HuH-7 cell line changed that. These cells are highly permissive to the replication of the hepatitis C virus, making them ideal for in vitro testing. By using the HuH-7 cells, researchers were able to screen drug candidates against laboratory-grown hepatitis C, which paved the way for the development of new drugs to fight the virus. Unlike other established human hepatoma cell lines, HuH-7 cells can be propagated in a chemically defined medium containing trace amounts of selenium in place of serum. This allows for systematic studies of the in vitro effects of various compounds on their growth and metabolism. |
|---|---|
| Organism | Human |
| Tissue | Liver |
| Disease | Hepatocellular carcinoma |
| Metastatic site | Hepatoma |
| Synonyms | HuH-7, HUH-7, Huh-7, Huh7, HUH7, HUH7.0, JTC-39, Japanese Tissue Culture-39 |
Details
| Age | 57 years |
|---|---|
| Gender | Male |
| Ethnicity | Japanese |
| Morphology | Epithelial-like |
| Growth properties | Adherent |
Documentation about the human hepatoma cell line HuH-7
| Citation | HuH7 (Cytion catalog number 300156) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_0336 |
Gene expression of HuH-7 cells
| Tumorigenic | Yes, in nude mice. |
|---|---|
| Viruses | Negative for HPV, HCV and HIV. |
Handling
| Culture Medium | RPMI 1640, w: 2.0 mM stable Glutamine, w: 2.0 g/L NaHCO3 (Cytion article number 820700a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Doubling time | 48 hours |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 1 to 2 x 104 cells/cm2 during routine cell culture |
| Fluid renewal | Every 3 days |
| Post-Thaw Recovery | Start culture using 2 to 3 x 104 cells/cm2. The cells will recover within 24 to 48 hours. |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality control of the HuH7 cell line
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300156-080724 | Certificate of Analysis | 23. May. 2025 | 300156 |
