HBL-100 Cells













USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
General information
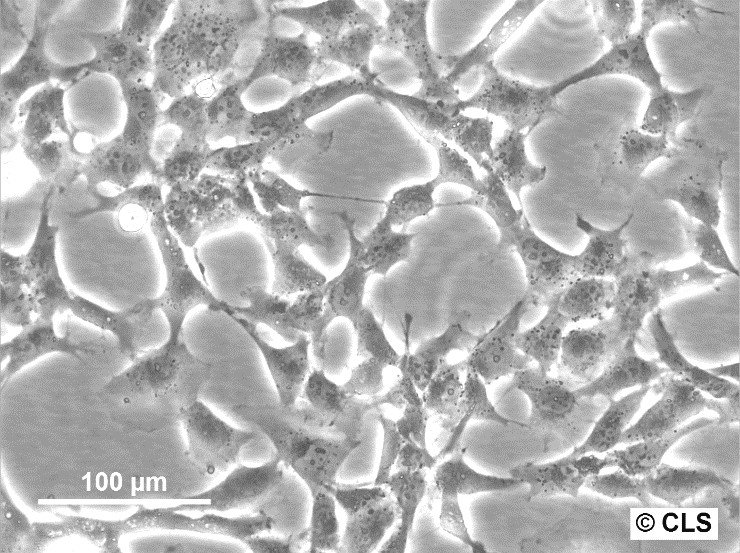
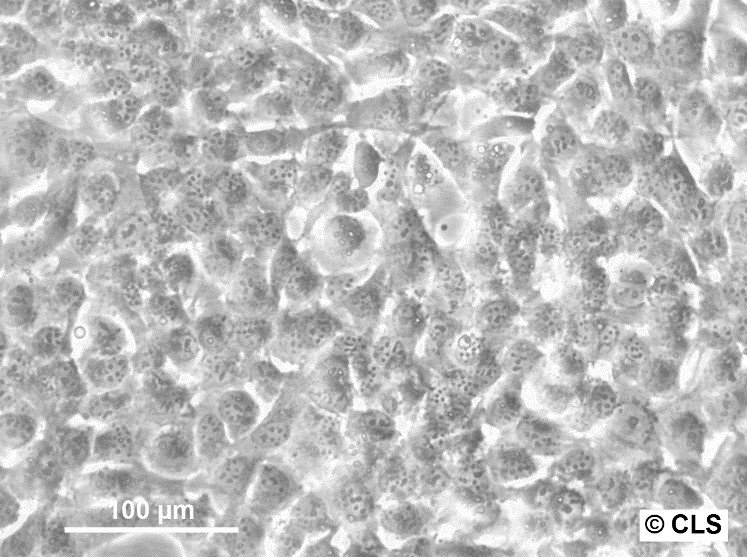
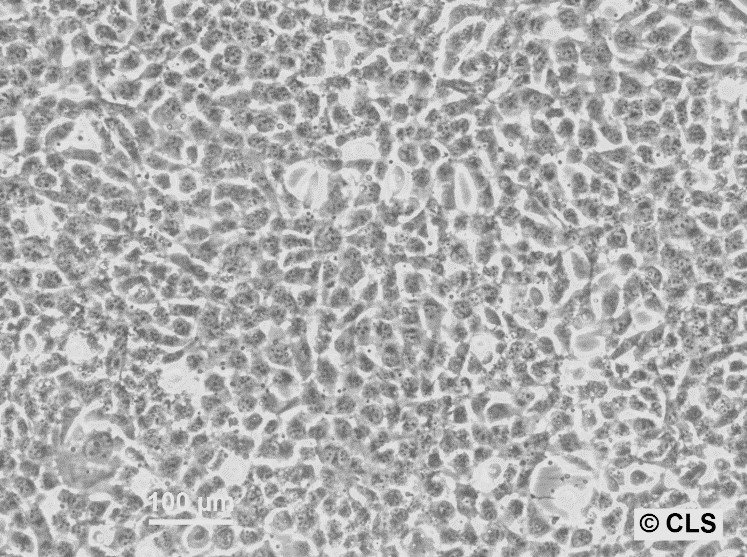
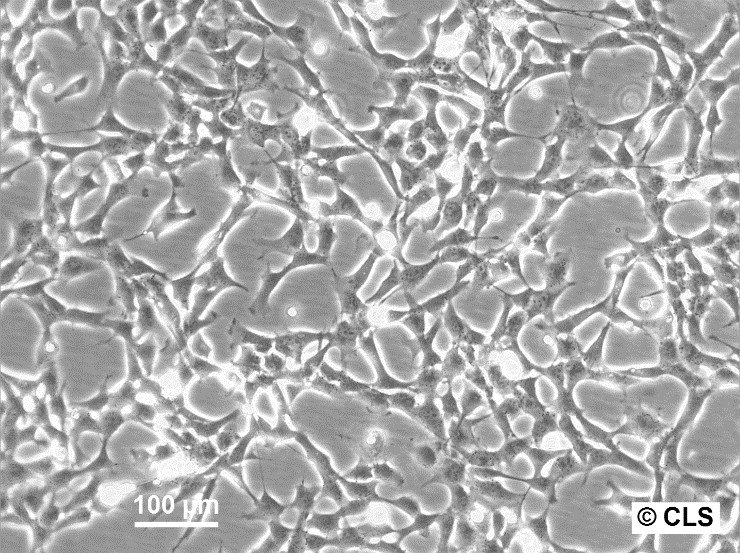
| Description | HBL-100 is a human breast epithelial cell line originally derived from the breast milk of a nursing mother. The milk was collected three days post-delivery, and despite no evidence of a breast lesion in the donor and no family history of breast cancer, the cells exhibited an abnormal karyotype by passage 7. This cell line is notable for its ability to synthesize a small amount of lactose and to respond to prolactin or estrogen stimulation by increasing the production of casein. Microscopic analyses, such as electron micrographs, have confirmed the presence of microvilli, tonofibrils, and desmosomes in these cells, highlighting their typical epithelial characteristics. However, the HBL-100 cell line has encountered significant complications regarding its identification and characterization. It was found to contain a Y chromosome, suggesting a misidentification as the cell line was initially thought to be of female origin. Further complexity arises from the presence of SV40 genomic sequences within the cell line, contradicting earlier beliefs that it was spontaneously immortalized. These findings have led to debates regarding the origin and the genetic makeup of HBL-100, making it a problematic cell line for research without thorough validation of its characteristics and origin. |
|---|---|
| Organism | Human |
| Tissue | Breast |
| Disease | Carcinoma |
| Synonyms | HBL 100, HBL100 |
Characteristics
| Age | 27 years |
|---|---|
| Gender | Female |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Monolayer, adherent |
Regulatory Data
| Citation | HBL-100 (Cytion catalog number 300178) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_4362 |
Biomolecular Data
| Antigen expression | HLA A1, A10, A11, B7, B8 |
|---|---|
| Isoenzymes | G6PD, B, PGM1, 1, PGM3, 2, ES-D, 1, Me-2, 0, GLO-1, 2, AK-1, 1-2, Phenotype Frequency Product: 0.0008 |
| Tumorigenic | Yes, in nude mice. At passage levels below 35 the line is not tumorigenic in nude mice, but forms colonies in soft agar. Tumorigenicity has been reported to increase above passage 35. |
| Viruses | The cells contain a tamdemly integrated SV40 genome it has been reported that they may contain a type D retrovirus that is similar or identical to Mason-Pfizer monkey virus (MPMV). |
| Reverse transcriptase | Positive |
| Ploidy status | Aneuploid |
| MSI-status | Stable (MSS) |
| Karyotype | The stemline chromosome number is near triploid with the modal number of 67 chromosomes, and the 2S component occurring at 0.6%. Most chromosome complements consist of about 39 normal and 28 marker chromosomes. Markers such as 2q, 11q+, 11q, t(2q.12), t(2q.5q?), t(6p?.16), 16pt and many others are common to most metaphases. Normal chromosomes 11, 14, 15 and 16 are absent. 2, 12, 17 and 19 are monosomic, and the x is disomic. DNA profiling for amelogenin, a sex-chromosome-specific PCR assay that can distinguish x chromosome-specific products from Y chromosome-specific products revealed the presence of Y chromosomes in this cell line of putative female origin. Confirmation of the general findings was accomplished by QM staining, C-banding, and FISH, with a whole chromosome paint probe to the human Y chromosome. |
Handling
| Culture Medium | McCoys 5a, w: 3.0 g/L Glucose, w: stable Glutamine, w: 2.0 mM Sodium pyruvate, w: 2.2 g/L NaHCO3 (Cytion article number 820200a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 1 x 104 cells/cm2 |
| Fluid renewal | 2 to 3 times per week |
| Post-Thaw Recovery | After thawing, plate the cells at 5 x 104 cells/cm2 and allow the cells to recover from the freezing process and to adhere for at least 24 hours. |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality Control & Molecular Analysis
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300178-619 | Certificate of Analysis | 23. May. 2025 | 300178 |
