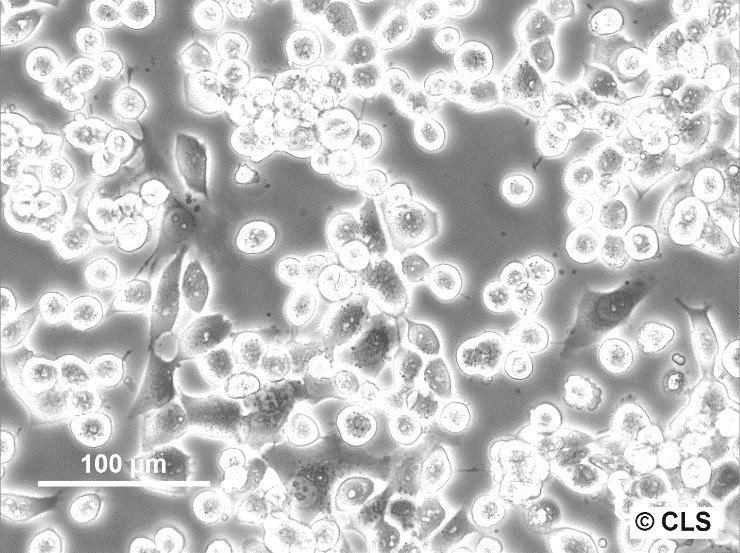
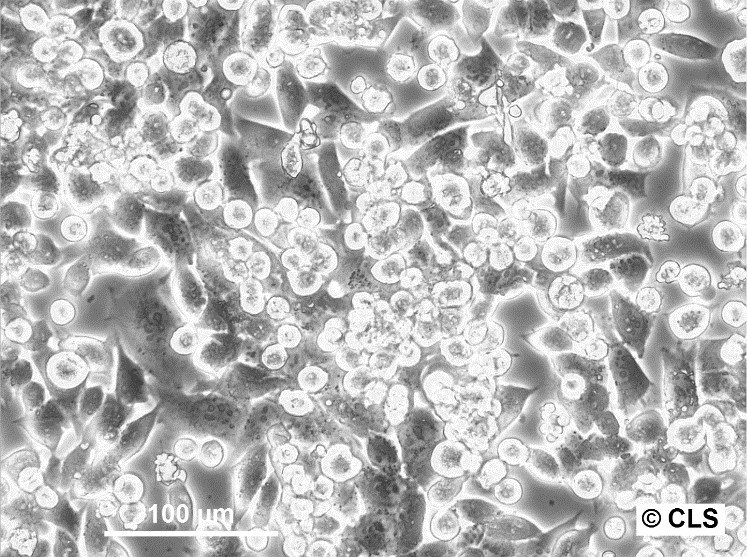
CERV-186 Cells














USD$800.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
General information
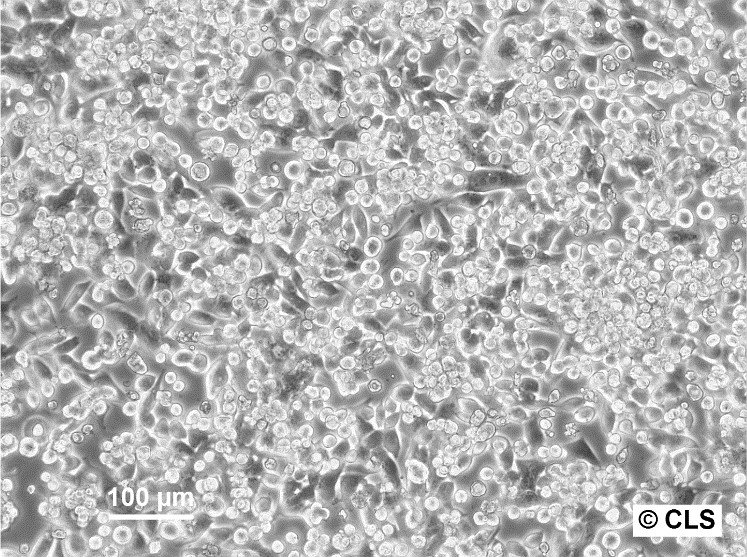
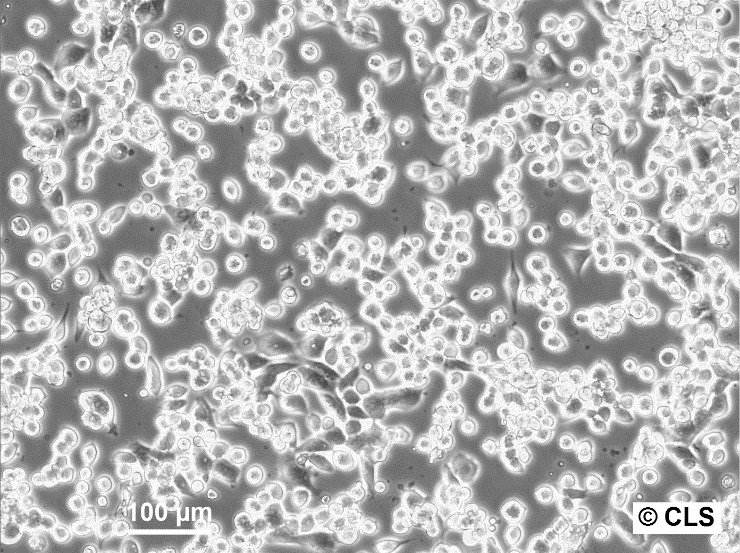
| Description | The CERV-186 cell line, derived in vitro from the xenotransplant of cervical carcinoma MRI-H-186, serves as a biological model for invasive, large-cell, non-keratinizing squamous cell carcinoma. This cell line was established and adapted for in vivo transplantation under the direction of Dr. Bodgen at the Mason Research Institute. Characterized by its genomic properties, MRI-H186 contains approximately 26 integrated copies of both full-length and truncated forms of the HPV16 genome, which significantly influence its transcriptomic profile. MRI-H186 cells are distinguished by their robust expression of both full-length and truncated early HPV16 transcripts, notably displaying high levels of E5 full-length (fl) RNA. This transcriptional signature is notably distinct from that observed in other cervical carcinoma cell lines such as CaSki and MRI-H196. Additionally, the transcriptional activity of MRI-H186, in terms of the expression of various other transcripts, shows a close alignment with the patterns observed in the HPK-IA and C3 cell lines, indicating a similar transcriptional behavior across these models. The presence of both full-length and truncated HPV16 genomic integrations in MRI-H186 cells is a key factor in their vigorous expression of early viral transcripts, particularly underscored by the significant expression of E5 fl RNA. This intense transcriptional activity culminates at the early polyadenylation signal, highlighting the unique transcriptional dynamics within the MRI-H186 cell line. |
|---|---|
| Organism | Human |
| Tissue | Cervix |
| Disease | Squamous cell carcinoma |
| Synonyms | Cerv-186, MRI-H-186, MRI-H186 |
Characteristics
| Age | 42 years |
|---|---|
| Gender | Female |
| Ethnicity | African |
| Morphology | Epithelial-like |
| Growth properties | Adherent |
Regulatory Data
| Citation | CERV-186 (Cytion catalog number 300290) |
|---|---|
| Biosafety level | 2 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_5720 |
Biomolecular Data
| Tumorigenic | Yes, in nude mice |
|---|---|
| Viruses | HPV-16 positive |
| Products | Cytokeratine 8, 18, Vimentin, Desmoplakin |
Handling
| Culture Medium | DMEM:Ham's F12 (1:1), w: 3.1 g/L Glucose, w: 2.5 mM L-Glutamine, w: 15 mM HEPES, w: 0.5 mM Sodium pyruvate, w: 1.2 g/L NaHCO3 (Cytion article number 820400a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 2 x 104 cells/cm2 will result in a confluent monolayer within 7 days |
| Fluid renewal | 2 to 3 times per week |
| Post-Thaw Recovery | After thawing, plate the cells at 5 x 104 cells/cm2 and allow the cells to recover from the freezing process and to adhere for at least 24 hours. |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality Control & Molecular Analysis
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300290-515SF | Certificate of Analysis | 23. May. 2025 | 300290 |
