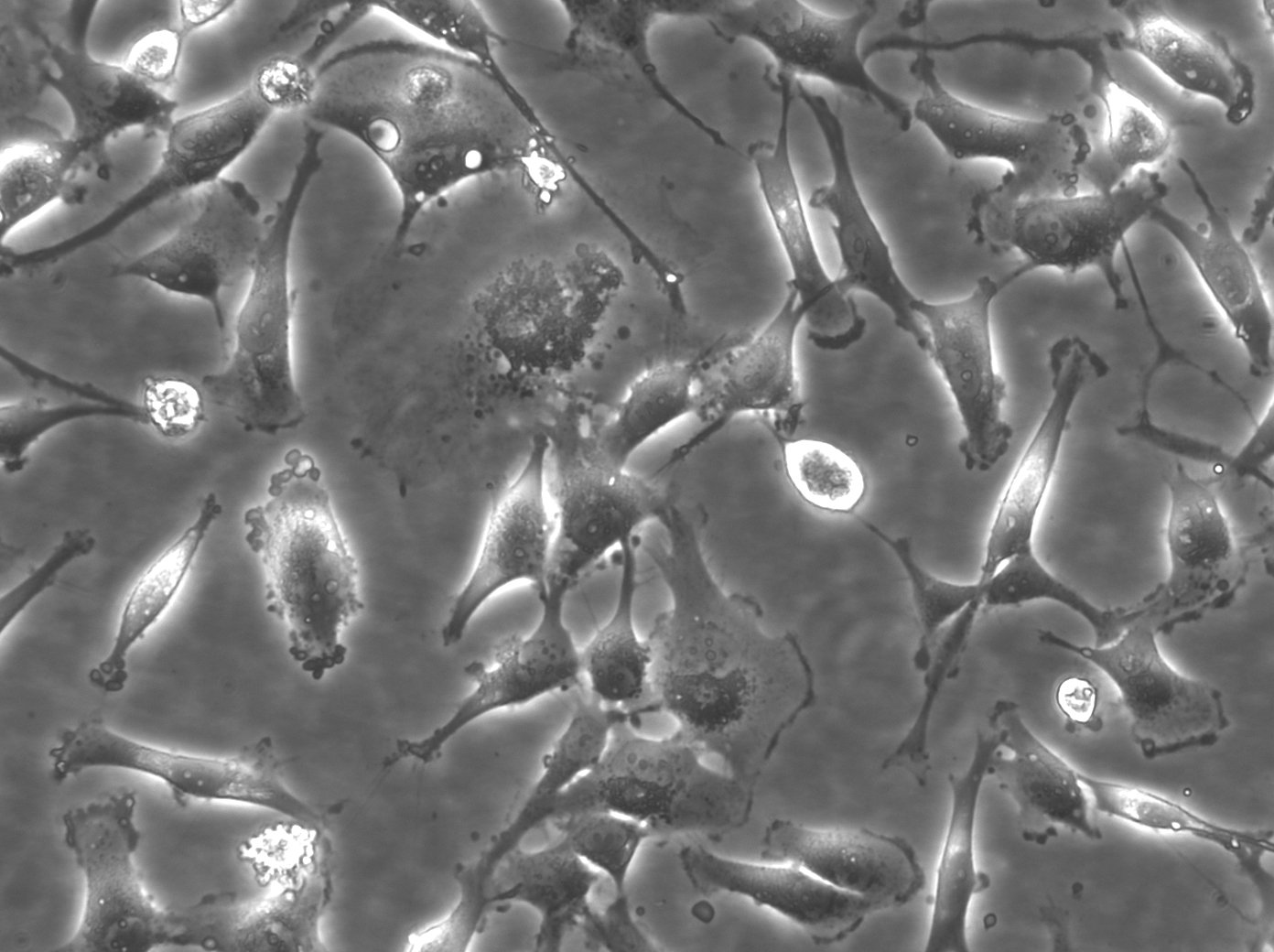
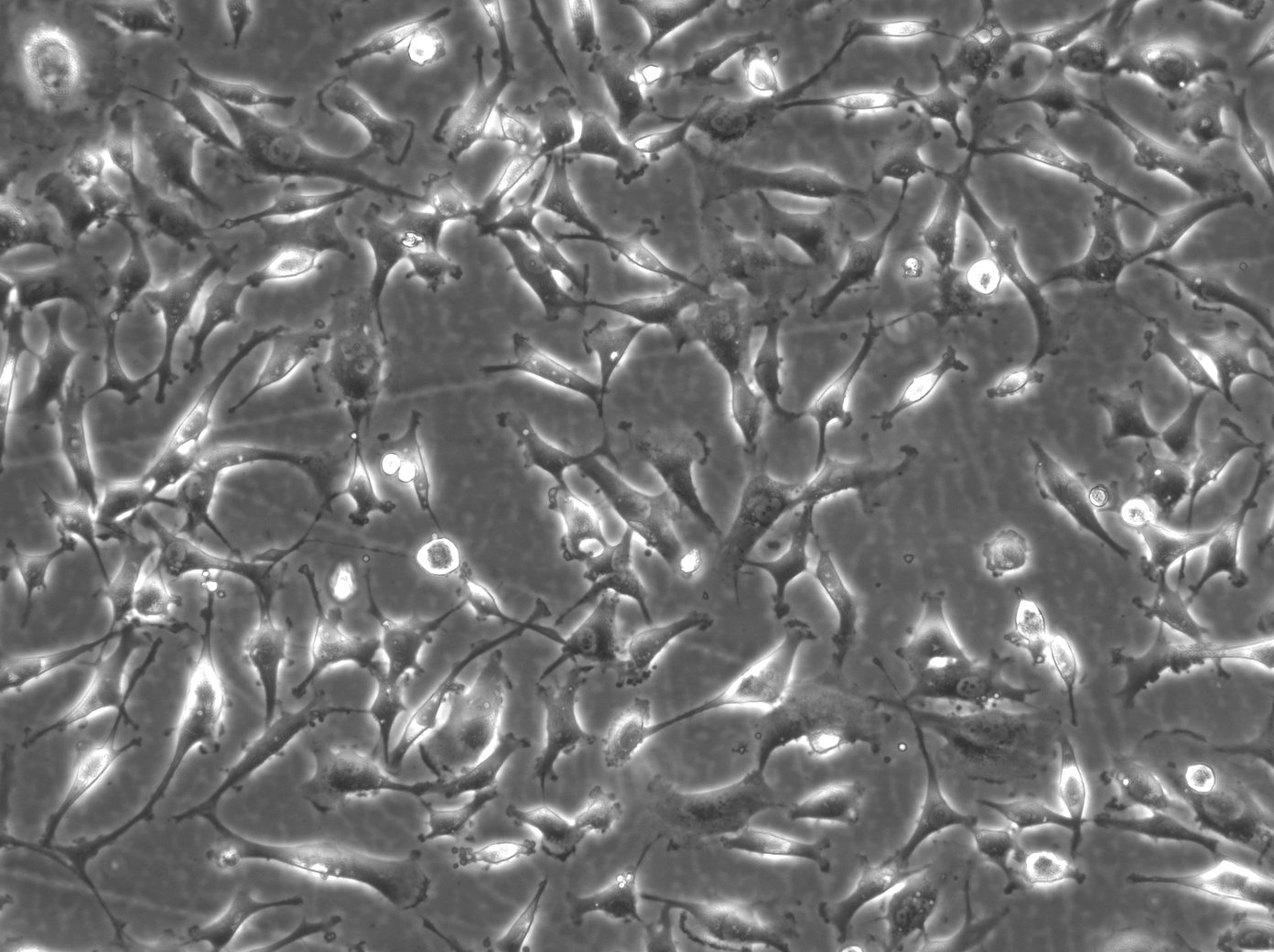
BT-549 Cells










USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
General information
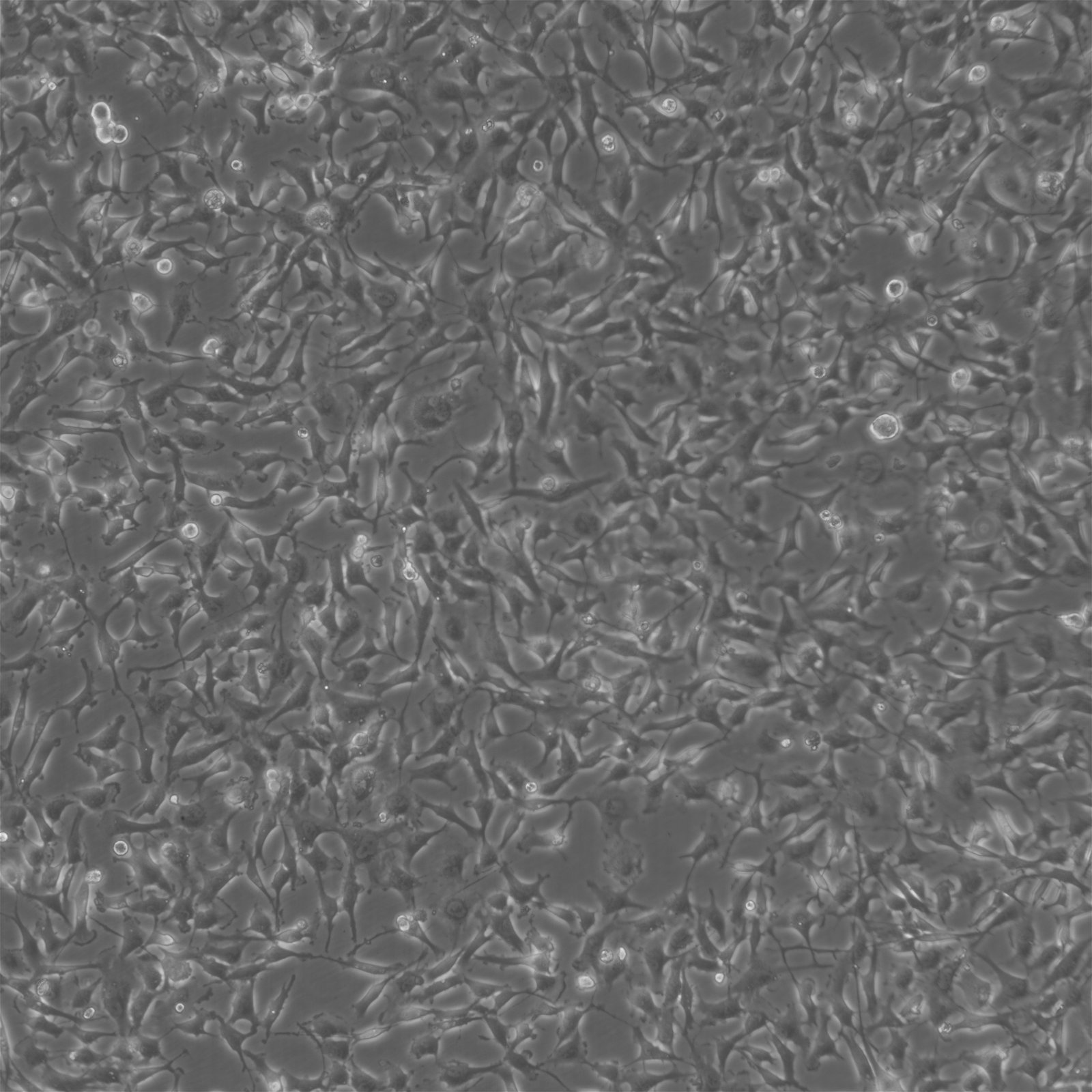
| Description | BT-549 cells are a human breast cancer cell line derived from the mammary gland tissue of a 72-year-old Caucasian woman with ductal carcinoma. They are commonly utilized in cancer research to study the biology and treatment of breast cancer, particularly the triple-negative subtype, which lacks estrogen receptor, progesterone receptor, and HER2 expression. BT-549 cells are characterized by their epithelial morphology and are known for their highly invasive properties, making them a valuable model for studying metastasis and tumor invasion. They exhibit several distinctive features including the presence of lipid droplets in the cytoplasm and a robust expression of the mucin-1 protein. These cells also express various oncogenes and tumor suppressor genes that are relevant to breast cancer pathology, such as TP53 and RB1. The BT-549 cell line is estrogen receptor-negative, progesterone receptor-negative, and does not amplify HER2, thus categorizing it under the triple-negative breast cancer (TNBC) subtype. Due to this classification, BT-549 cells are particularly useful for studying the unique mechanisms of progression and treatment response in TNBC, which is known for its aggressive nature and lack of targeted therapies. Furthermore, BT-549 cells are often used in drug resistance studies and for testing new chemotherapeutic agents and targeted therapies, offering insights into potential therapeutic strategies for managing and treating aggressive forms of breast cancer. |
|---|---|
| Organism | Human |
| Tissue | Breast, mammary gland |
| Disease | Invasive ductal carcinoma |
| Metastatic site | Ductal |
| Synonyms | BT 549, BT.549, BT549 |
Characteristics
| Age | 72 years |
|---|---|
| Gender | Female |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Monolayer, adherent |
Regulatory Data
| Citation | BT-549 (Cytion catalog number 300132) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_1092 |
Biomolecular Data
| Isoenzymes | G6PD, B, PGM1, 2, PGM3, 1, ES-D, 1, Me-2, 1, AK-1, 1, GLO-1, 1-2, Phenotype Frequency Product: 0.0048 |
|---|---|
| Mutational profile | TP53 mut |
| Karyotype | Mode = 74, range = 53 to 140, three marker chromosomes |
Handling
| Culture Medium | DMEM, w: 4.5 g/L Glucose, w: 4 mM L-Glutamine, w: 3.7 g/L NaHCO3, w: 1.0 mM Sodium pyruvate (Cytion article number 820300a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 1 x 104 cells/cm2 will yield in a confluent layer in about 4 days |
| Fluid renewal | 2 to 3 times per week |
| Post-Thaw Recovery | After thawing, plate the cells at 5 x 104 cells/cm2 and allow the cells to recover from the freezing process and to adhere for at least 24 hours. |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality Control & Molecular Analysis
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300132-1121SF | Certificate of Analysis | 23. May. 2025 | 300132 |
| 300132-310724 | Certificate of Analysis | 23. May. 2025 | 300132 |
