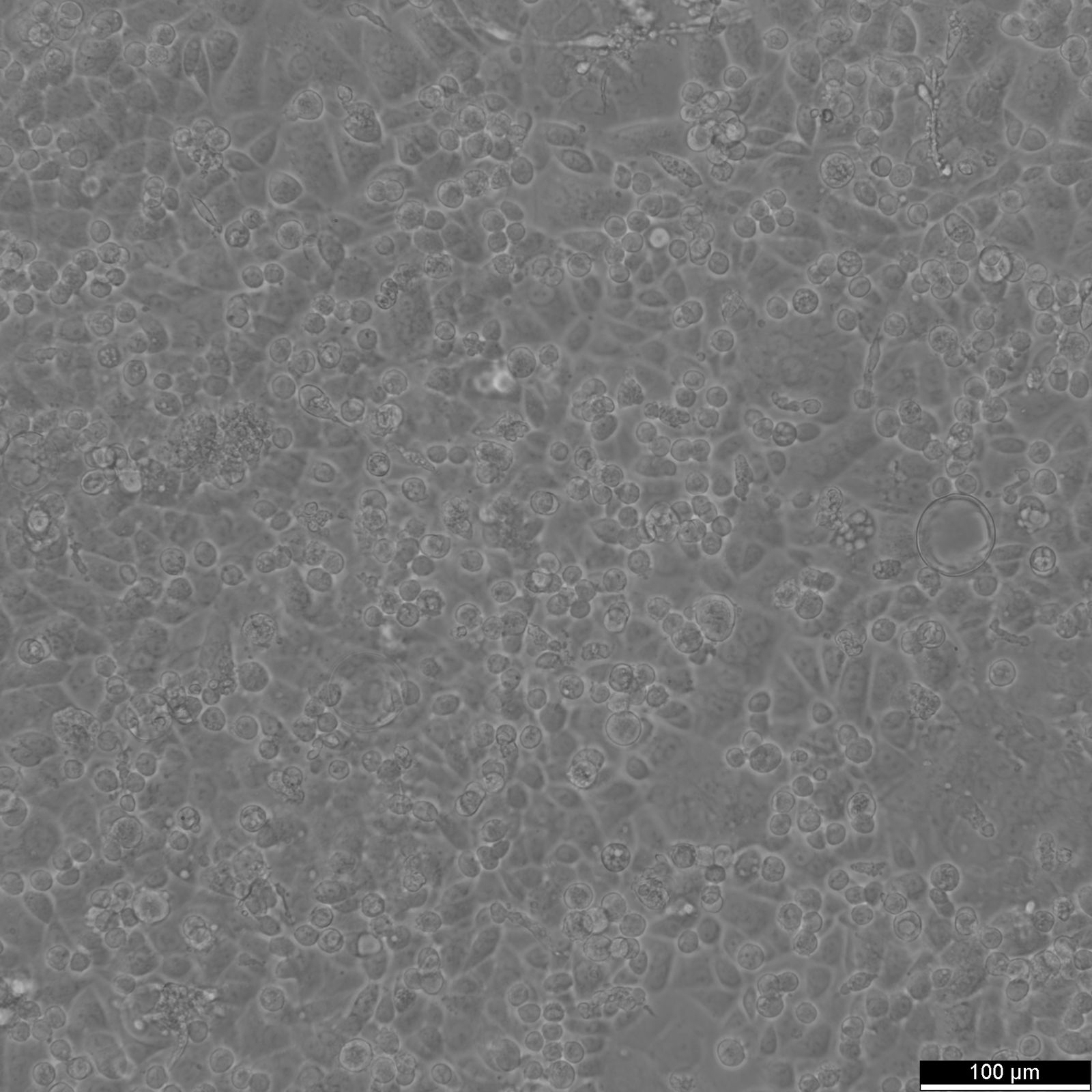
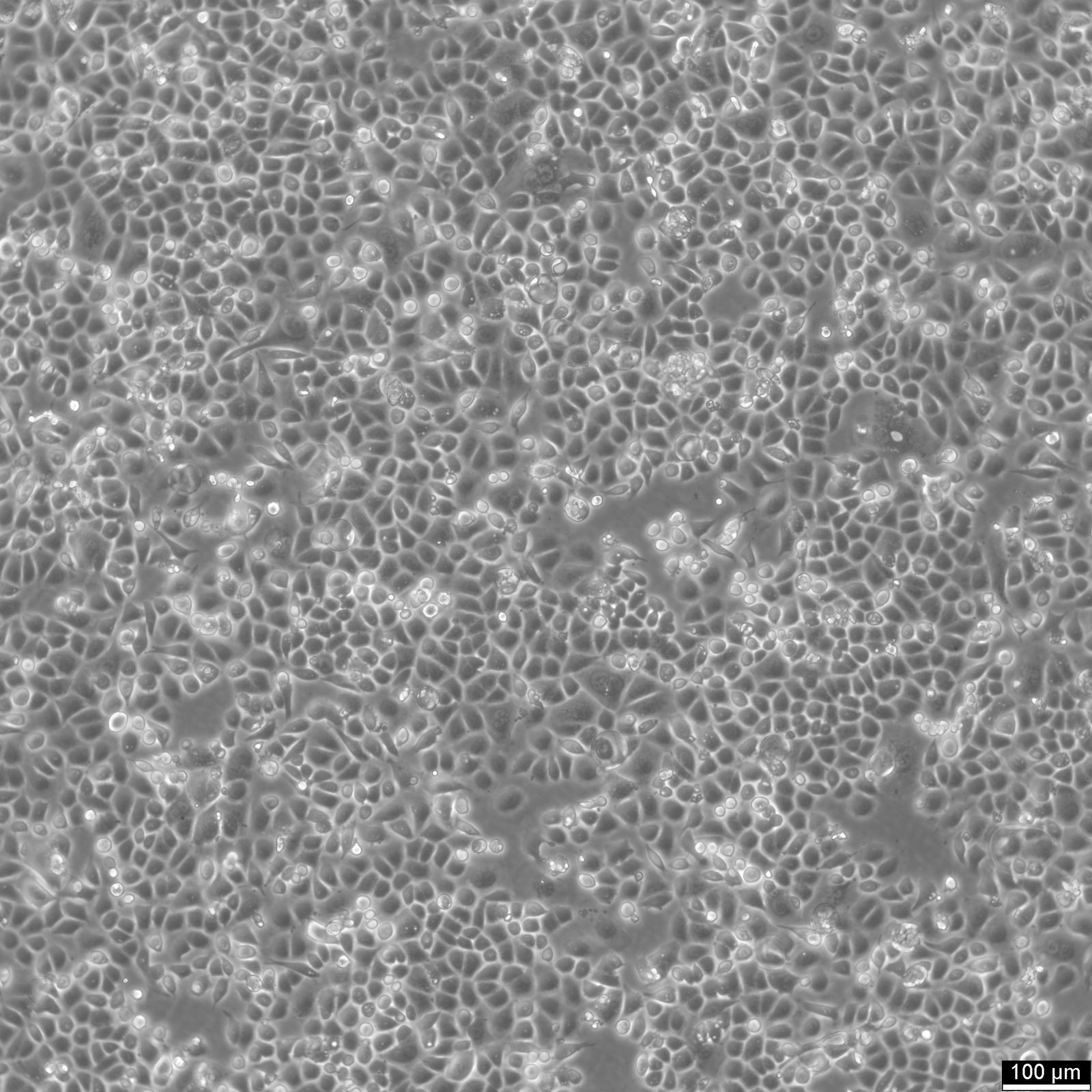
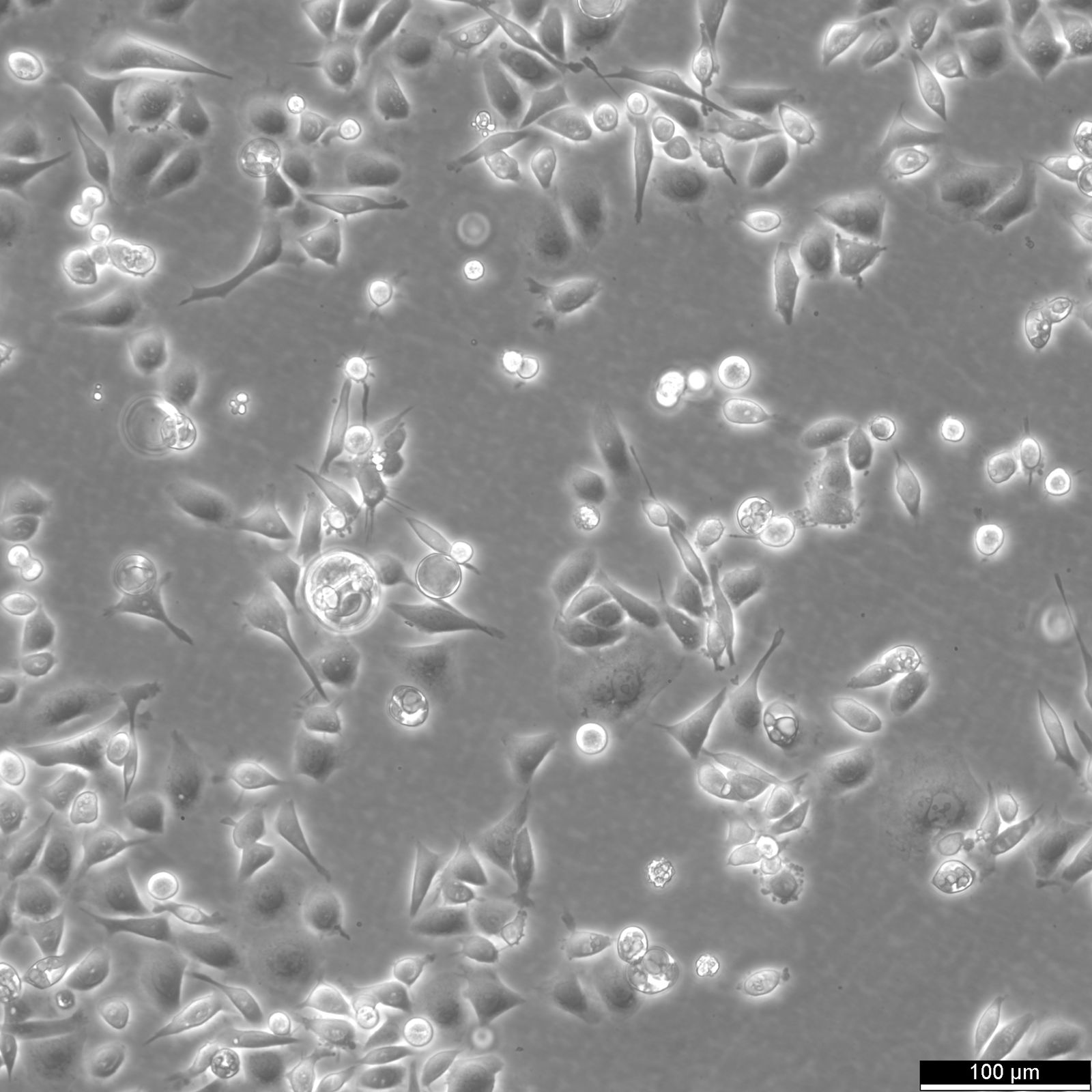
AGS Cells



















USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
General information
| Description | AGS cells are a human gastric adenocarcinoma cell line derived from the stomach tissue of a 54-year-old Caucasian female. They are extensively used in biomedical research focused on gastric cancer, including studies on cancer cell biology, pathogenesis, and drug testing. The AGS cell line exhibits epithelial-like morphology and is characterized by its aggressive growth pattern and tumorigenic potential in vivo. These cells are commonly used as a model to study the molecular and cellular mechanisms underlying gastric carcinogenesis, including the influence of Helicobacter pylori infection, a well-known risk factor for gastric cancer. AGS cells provide a robust system to explore the interactions between gastric cancer cells and H. pylori, especially regarding how bacterial factors affect cancer cell proliferation, apoptosis, and inflammatory responses. AGS cells are also valuable for examining the gastric epithelial barrier's response to various stimuli, including inflammatory cytokines, and for studying signaling pathways implicated in gastric cancer, such as those involving NF-kB, Wnt, and MAPK. Their utility extends to the assessment of new therapeutic agents, where they are used to evaluate the efficacy and mechanisms of action of anticancer drugs, targeted therapies, and natural compounds with potential anti-cancer properties. Furthermore, AGS cells are often employed in studies aimed at understanding the genetic and epigenetic alterations in gastric cancer, offering insights into potential diagnostic markers and therapeutic targets for this challenging and frequently fatal disease. |
|---|---|
| Organism | Human |
| Tissue | Gastric |
| Disease | Adenocarcinoma |
Characteristics
| Age | 54 years |
|---|---|
| Gender | Female |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Monolayer, adherent |
Regulatory Data
| Citation | AGS (Cytion catalog number 300408) |
|---|---|
| Biosafety level | 2 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_0139 |
Biomolecular Data
| Protein expression | P53 positive |
|---|---|
| Tumorigenic | Yes, in athymic BALB/c mice |
| Viruses | This cell line may release Parainfluenzavirus Type 5 (formerly known as Simian Virus 5). The virus interferes with Interferon-signalling within the cell line by degradation of STAT1. |
| Karyotype | Modal number = 47, range = 39 to 92 |
Handling
| Culture Medium | DMEM, w: 4.5 g/L Glucose, w: 4 mM L-Glutamine, w: 3.7 g/L NaHCO3, w: 1.0 mM Sodium pyruvate (Cytion article number 820300a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Doubling time | 24 to 48 hours |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 1 x 104 cells/cm2 will result in a confluent monolayer within 3to5 days. |
| Fluid renewal | 2 to 3 times per week |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality Control & Molecular Analysis
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300408-814 | Certificate of Analysis | 23. May. 2025 | 300408 |
| 300408-220425 | Certificate of Analysis | 23. May. 2025 | 300408 |
| 300408-220124 | Certificate of Analysis | 05. Dec. 2025 | 300408 |
