769-P Cells
















USD$395.00*
Products are shipped frozen on dry ice in cryotubes. Each cryotube typically contains 3 × 106 cells for adherent lines or 5 × 106 cells for suspension lines (refer to the batch CoA for details).
General information
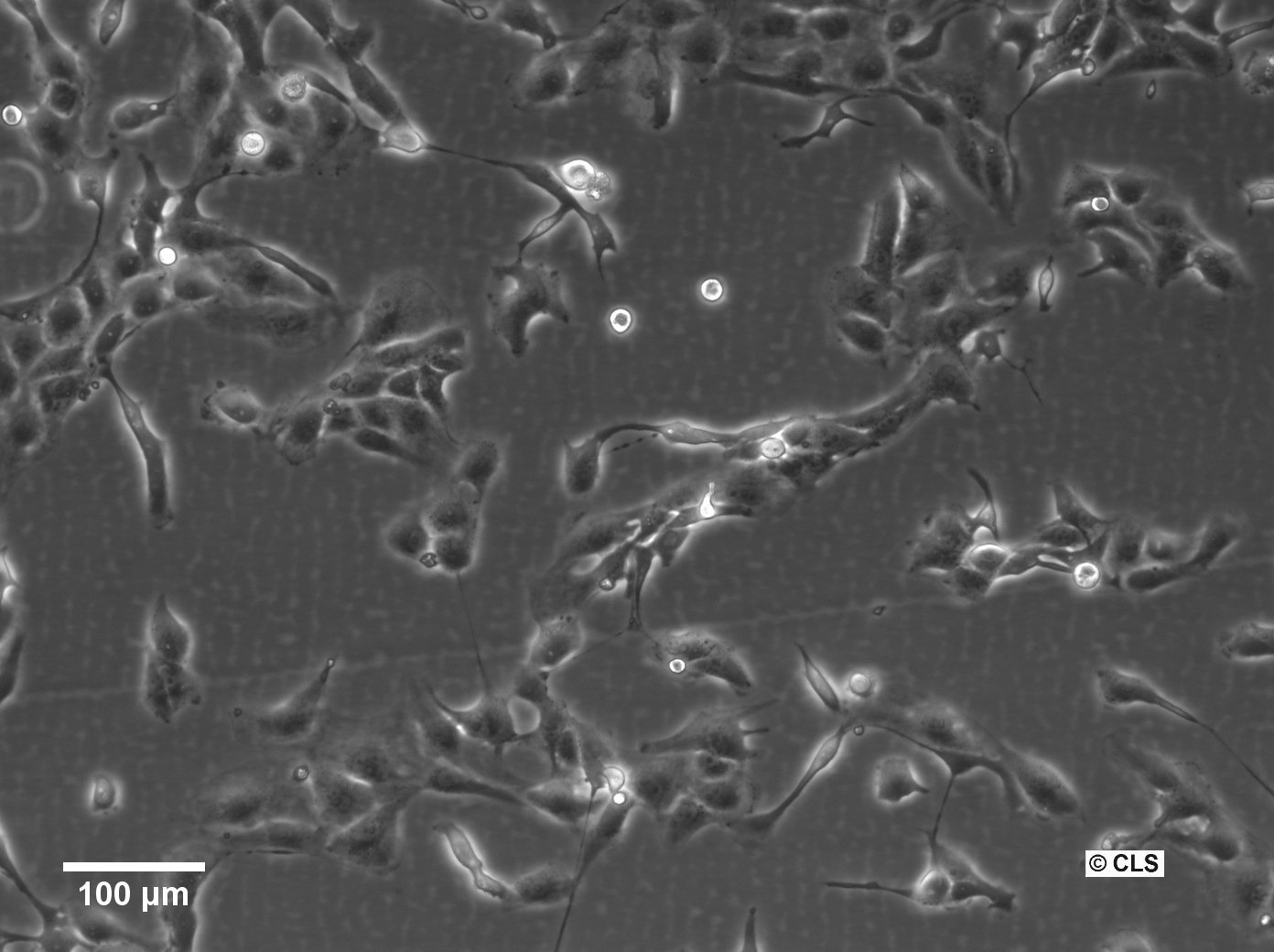
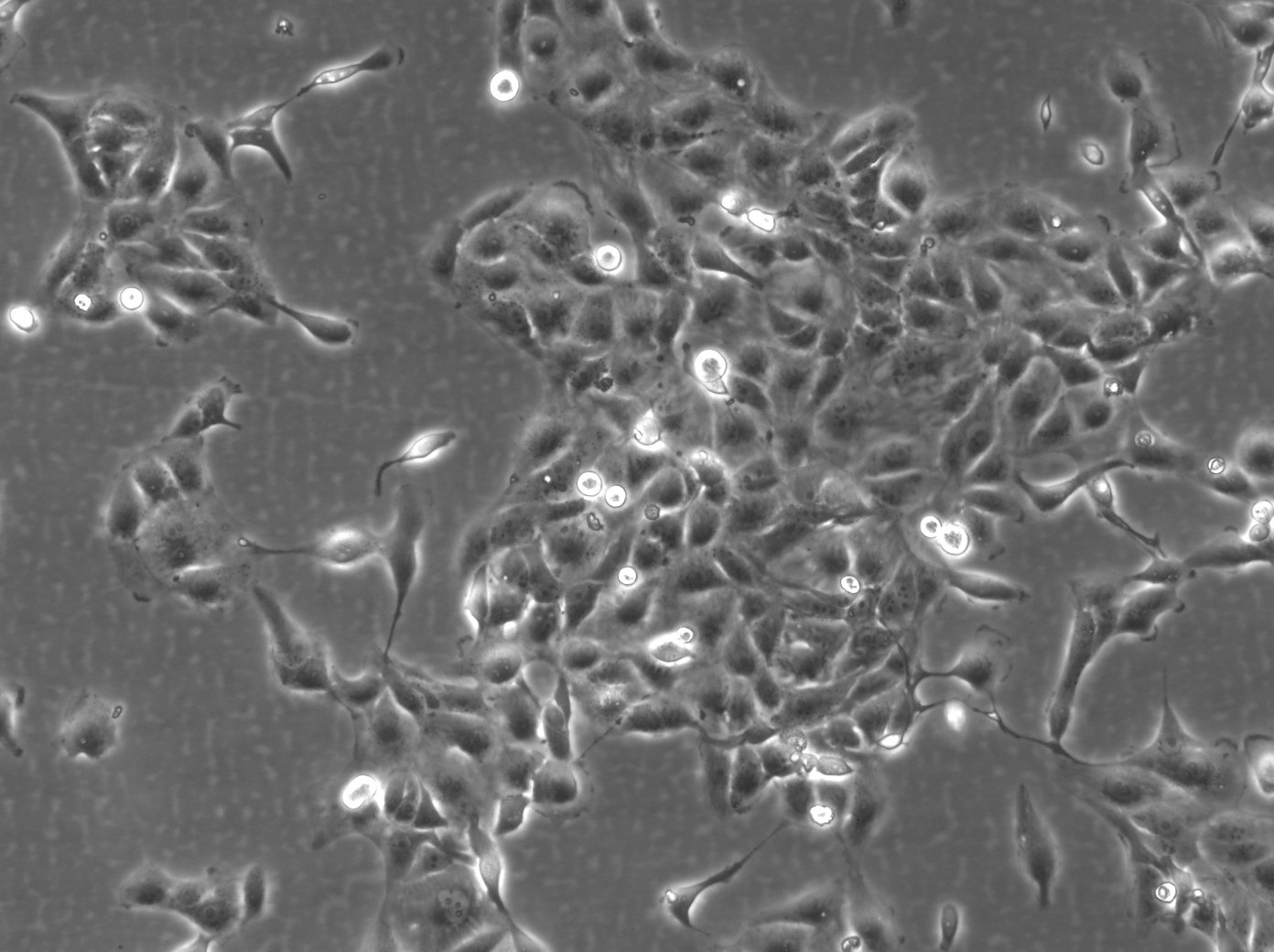
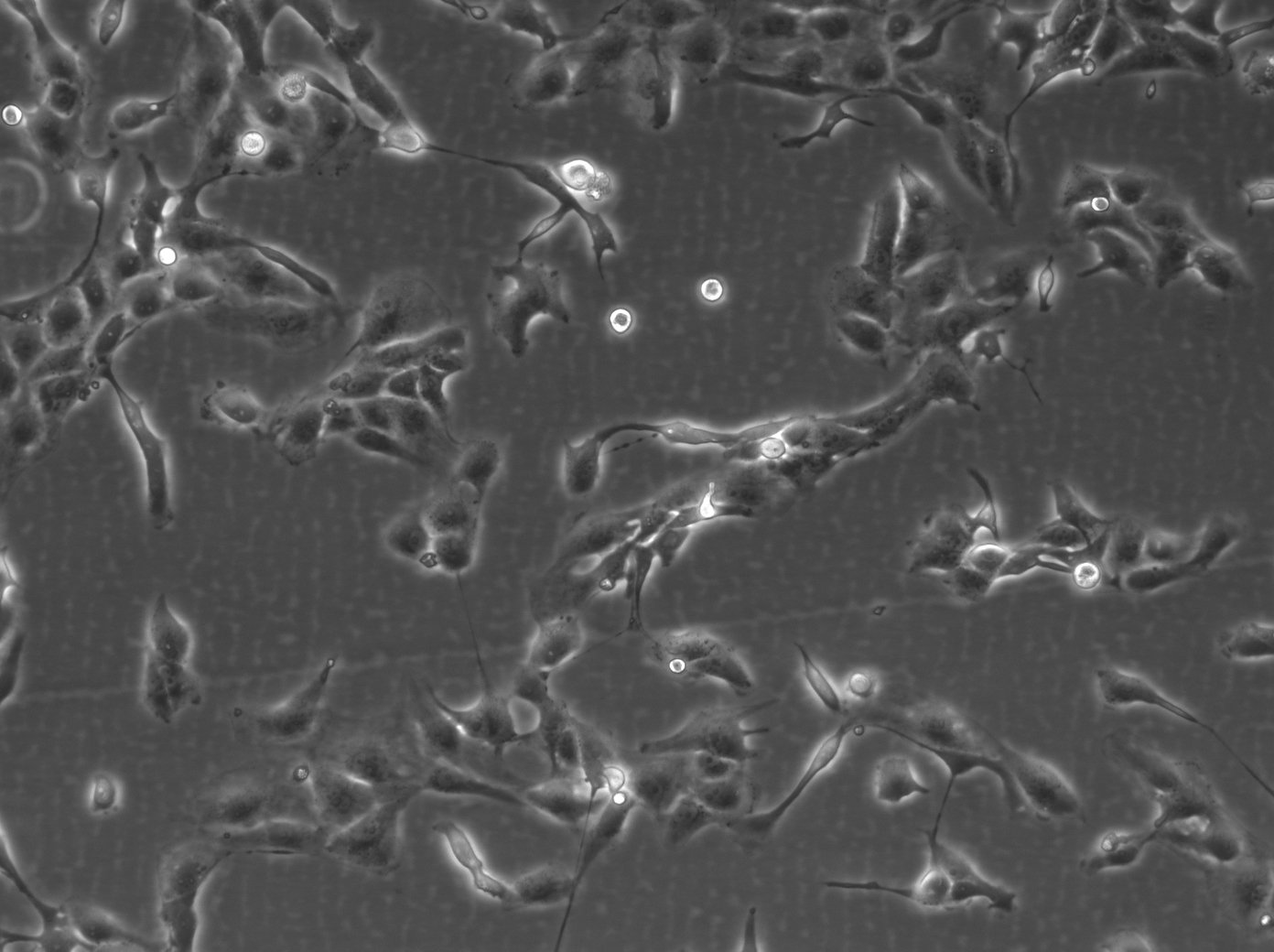
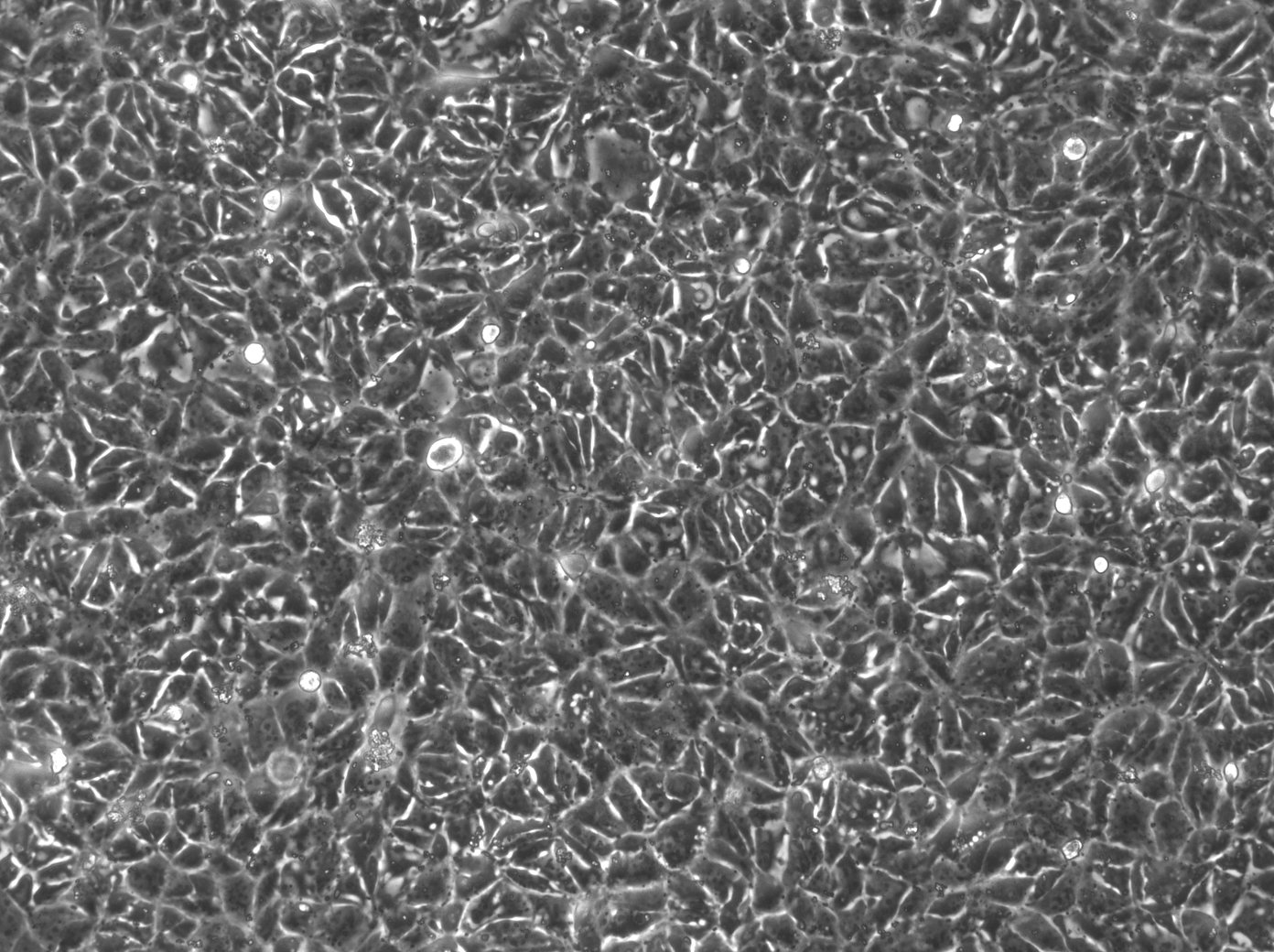
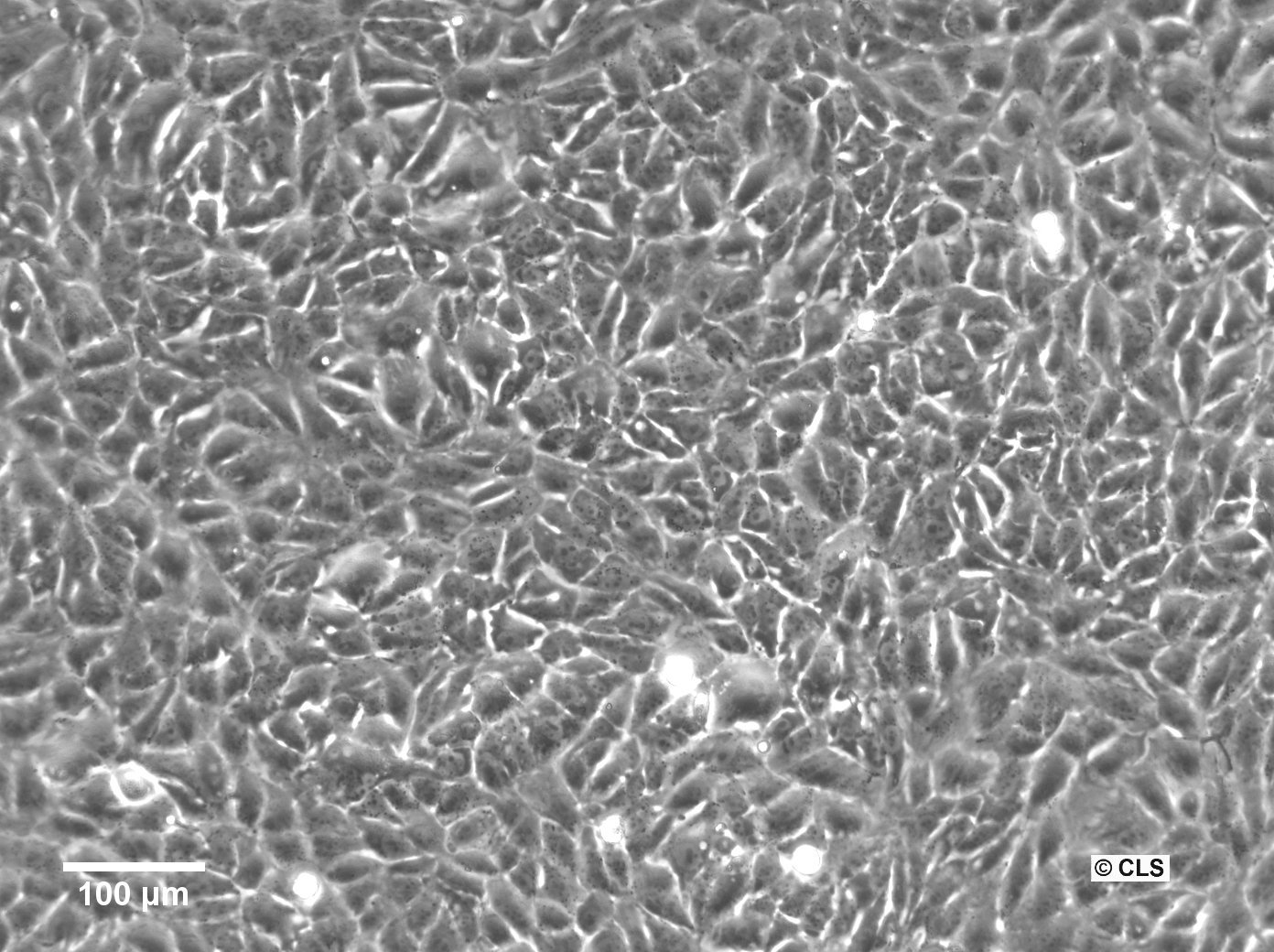
| Description | The 769-P cell line is a human renal cell carcinoma (RCC) cell line that was derived from a nephrectomy specimen of a 63-year-old female patient with renal cell adenocarcinoma in 1975. It is widely used in renal cell cancer research, particularly clear cell renal cell carcinoma (ccRCC), which is the most common and lethal form of kidney cancer in adults. The 769-P cell line retains many characteristics of primary RCC and harbors several mutations that are relevant to renal cell carcinoma. They exhibit a loss of function in the von Hippel-Lindau (VHL) tumor suppressor gene, which is an important kidney cancer gene in ccRCC that can activate various oncogenic pathways including angiogenesis, cell proliferation, and metabolic reprogramming. The 769-P cell line is used to understand the molecular mechanisms of kidney cancer pathogenesis, explore the efficacy of anticancer drugs, and investigate the mechanisms of drug resistance. These cells are particularly useful for studying the response to tyrosine kinase inhibitors (TKIs), which are a class of targeted therapies used in the treatment of RCC and RCC subtypes. The 769-P renal cancer cell line is further used to investigate the role of the tumor microenvironment in kidney cancer and to study cellular processes like apoptosis, cell cycle regulation, and metastatic potential. Their responsiveness to hypoxic conditions makes them suitable for research into how ccRCC adapts to and thrives in low-oxygen environments found within solid tumors. In summary, the 769-P cell line and other RCC cell lines are indispensable tools in renal carcinoma research, providing insights into the pathogenesis of ccRCC, drug efficacy, and resistance mechanisms. |
|---|---|
| Organism | Human |
| Tissue | Kidney |
| Disease | Renal cell carcinoma |
| Synonyms | 769P, 769-p |
Characteristics
| Age | 63 years |
|---|---|
| Gender | Female |
| Ethnicity | Caucasian |
| Morphology | Epithelial-like |
| Growth properties | Monolayer, adherent |
Regulatory Data
| Citation | 769-P (Cytion catalog number 300106) |
|---|---|
| Biosafety level | 1 |
| NCBI_TaxID | 9606 |
| CellosaurusAccession | CVCL_1050 |
Biomolecular Data
| Tumorigenic | Forms tumors in immunosuppressed hamsters and in nude mice |
|---|---|
| Ploidy status | This cell line had a high number of tetra-, hexa-, and higher-ploid cells (2s populations). The most common cell population (32% of cells) had a pseudodiploid karyotype of 46,xx,-3,-18,del(7) (q21.12,q22.3), ?t(3q?18q). |
| Karyotype | Hypodiploid. Modal number = 45. A large submetacentric chromosome was present in all cells. |
Handling
| Culture Medium | RPMI 1640, w: 2.0 mM stable Glutamine, w: 2.0 g/L NaHCO3 (Cytion article number 820700a) |
|---|---|
| Supplements | Supplement the medium with 10% FBS |
| Dissociation Reagent | Accutase |
| Doubling time | 35 hours |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Seeding density | 3 x 104 cells/cm2 will result in a confluent monolayer within 4 days. |
| Fluid renewal | 2 to 3 times per week |
| Post-Thaw Recovery | After thawing, plate the cells at 5 x 104 cells/cm2 and allow the cells to recover from the freezing process and to adhere for at least 48 hours. |
| Freeze medium | As a cryopreservation medium, we use complete growth medium (including FBS) + 10% DMSO for adequate post-thaw viability, or CM-1 (Cytion catalog number 800100), which includes optimized osmoprotectants and metabolic stabilizers to enhance recovery and reduce cryo-induced stress. |
| Thawing and Culturing Cells |
|
| Incubation Atmosphere | 37°C, 5% CO2, humidified atmosphere. |
| Shipping Conditions | Cryopreserved cell lines are shipped on dry ice in validated, insulated packaging with sufficient refrigerant to maintain approximately −78 °C throughout transit. On receipt, inspect the container immediately and transfer vials without delay to appropriate storage. |
| Storage Conditions | For long-term preservation, place vials in vapor-phase liquid nitrogen at about −150 to −196 °C. Storage at −80 °C is acceptable only as a short interim step before transfer to liquid nitrogen. |
Quality Control & Molecular Analysis
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|
Certificate of Analysis (CoA)
| Lot Number | Certificate Type | Date | Catalog Number |
|---|---|---|---|
| 300106-721 | Certificate of Analysis | 23. May. 2025 | 300106 |
